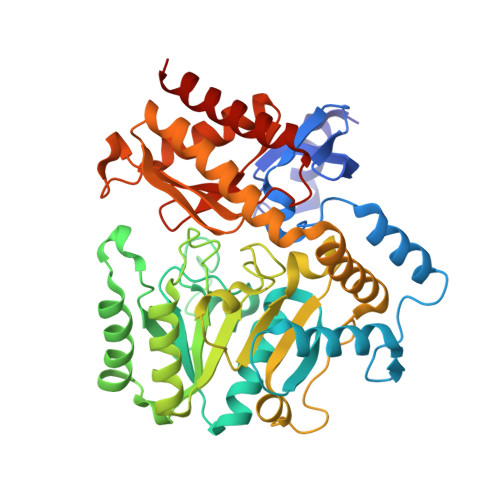
Crystal structures of unbound and aminooxyacetate-bound Escherichia coli gamma-aminobutyrate aminotransferase.
Liu, W., Peterson, P.E., Carter, R.J., Zhou, X., Langston, J.A., Fisher, A.J., Toney, M.D.(2004) Biochemistry 43: 10896-10905
- PubMed: 15323550 Search on PubMed
- DOI: https://doi.org/10.1021/bi049218e
- Primary Citation Related Structures:
1SF2, 1SFF - PubMed Abstract:
The X-ray crystal structures of Escherichia coli gamma-aminobutyrate aminotransferase unbound and bound to the inhibitor aminooxyacetate are reported. The enzyme crystallizes from ammonium sulfate solutions in the P3(2)21 space group with a tetramer in the asymmetric unit. Diffraction data were collected to 2.4 A resolution for the unliganded enzyme and 1.9 A resolution for the aminooxyacetate complex. The overall structure of the enzyme is similar to those of other aminotransferase subgroup II enzymes. The ability of gamma-aminobutyrate aminotransferase to act on primary amine substrates (gamma-aminobutyrate) in the first half-reaction and alpha-amino acids in the second is proposed to be enabled by the presence of Glu211, whose side chain carboxylate alternates between interactions with Arg398 in the primary amine half-reaction and an alternative binding site in the alpha-amino acid half-reaction, in which Arg398 binds the substrate alpha-carboxylate. The specificity for a carboxylate group on the substrate side chain is due primarily to the presence of Arg141, but also requires substantial local main chain rearrangements relative to the structurally homologous enzyme dialkylglycine decarboxylase, which is specific for small alkyl side chains. No iron-sulfur cluster is found in the bacterial enzyme as was found in the pig enzyme [Storici, P., De Biase, D., Bossa, F., Bruno, S., Mozzarelli, A., Peneff, C., Silverman, R. B., and Schirmer, T. (2004) J. Biol. Chem. 279, 363-73.]. The binding of aminooxyacetate causes remarkably small changes in the active site structure, and no large domain movements are observed. Active site structure comparisons with pig gamma-aminobutyrate aminotransferase and dialkylglycine decarboxylase are discussed.
- Department of Chemistry, University of California, Davis, California 95616, USA.
Organizational Affiliation: