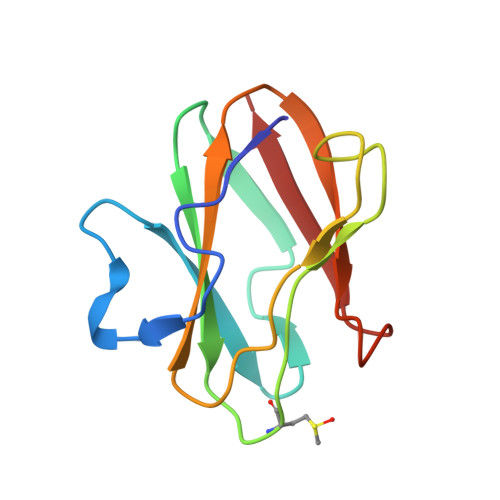
Structural Studies of Two Mutants of Amicyanin from Paracoccus denitrificans That Stabilize the Reduced State of the Copper.
Carrell, C.J., Sun, D., Jiang, S., Davidson, V.L., Mathews, F.S.(2004) Biochemistry 43: 9372-9380
- PubMed: 15260480 Search on PubMed
- DOI: https://doi.org/10.1021/bi049634z
- Primary Citation Related Structures:
1SF3, 1SF5, 1SFD, 1SFH - PubMed Abstract:
Mutation of Pro94 to phenylalanine or alanine significantly alters the redox properties of the type I copper center of amicyanin. Each mutation increases the redox midpoint potential (E(m)) value by at least 140 mV and shifts the pK(a) for the pH dependence of the E(m) value to a more acidic value. Atomic resolution (0.99-1.1 A) structures of both the P94F and P94A amicyanin have been determined in the oxidized and reduced states. In each amicyanin mutant, an electron-withdrawing hydrogen bond to the copper-coordinating thiolate sulfur of Cys92 is introduced by movement of the amide nitrogens of Phe94 and Ala94 much closer to the thiolate sulfur than in wild-type amicyanin. This is the likely explanation for the much more positive E(m) values which result from each of these mutations. The observed decrease in the pK(a) value for the pH dependence of the E(m) value that is seen in the mutants seems to be correlated with steric hindrance to the rotation of the His95 copper ligand which results from the mutations. In wild-type amicyanin the His95 side chain undergoes a redox and pH-dependent conformational change which accounts for the pH dependence of the E(m) value of amicyanin. The reduced P94A amicyanin exhibits two alternate conformations with the positions of the copper 1.4 A apart. In one of these conformations, a water molecule appears to have replaced Met98 as a copper ligand. The relevance of these structures to the electron transfer properties of P94F and P94A amicyanin are also discussed.
- Department of Biochemistry and Molecular Biophysics, Washington University School of Medicine, St. Louis, Missouri 63110, USA.
Organizational Affiliation: