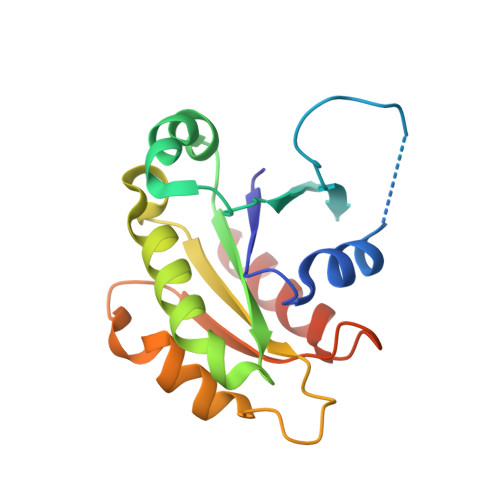
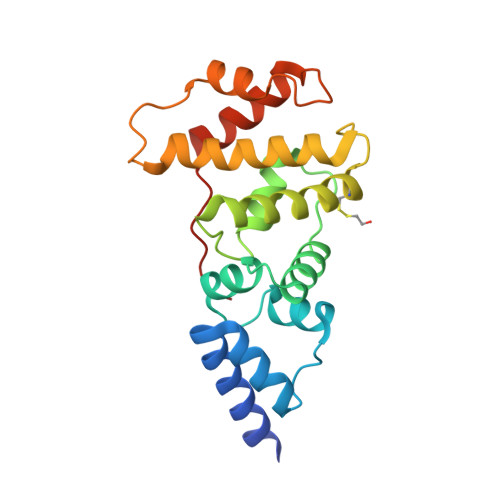
Structural Snapshots of the Mechanism and Inhibition of a Guanine Nucleotide Exchange Factor
Renault, L., Guibert, B., Cherfils, J.(2003) Nature 426: 525-530
- PubMed: 14654833 Search on PubMed
- DOI: https://doi.org/10.1038/nature02197
- Primary Citation Related Structures:
1R8M, 1R8Q, 1R8S, 1S9D - PubMed Abstract:
Small GTP-binding (G) proteins are activated by GDP/GTP nucleotide exchange stimulated by guanine nucleotide exchange factors (GEFs). Nucleotide dissociation from small G protein-GEF complexes involves transient GDP-bound intermediates whose structures have never been described. In the case of Arf proteins, small G proteins that regulate membrane traffic in eukaryotic cells, such intermediates can be trapped either by the natural inhibitor brefeldin A or by charge reversal at the catalytic glutamate of the Sec7 domain of their GEFs. Here we report the crystal structures of these intermediates that show that membrane recruitment of Arf and nucleotide dissociation are separate reactions stimulated by Sec7. The reactions proceed through sequential rotations of the Arf.GDP core towards the Sec7 catalytic site, and are blocked by interfacial binding of brefeldin A and unproductive stabilization of GDP by charge reversal. The structural characteristics of the reaction and its modes of inhibition reveal unexplored ways in which to inhibit the activation of small G proteins.
- Laboratoire d'Enzymologie et Biochimie Structurales, CNRS UPR 9063, Avenue de la Terrasse, 91198 Gif sur Yvette, France.
Organizational Affiliation: