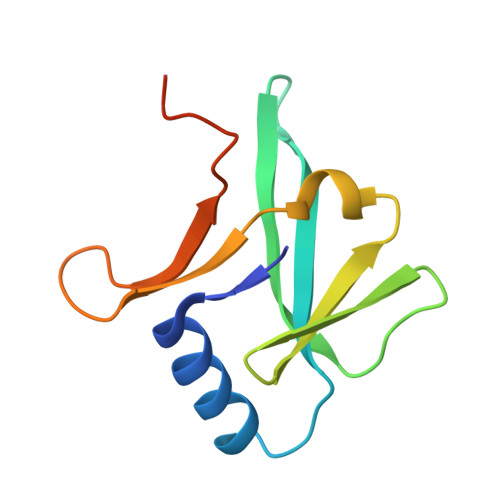
Crystal structure of IscA, an iron-sulfur cluster assembly protein from Escherichia coli.
Cupp-Vickery, J.R., Silberg, J.J., Ta, D.T., Vickery, L.E.(2004) J Mol Biology 338: 127-137
- PubMed: 15050828 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.02.027
- Primary Citation Related Structures:
1S98 - PubMed Abstract:
IscA, an 11 kDa member of the hesB family of proteins, binds iron and [2Fe-2S] clusters, and participates in the biosynthesis of iron-sulfur proteins. We report the crystal structure of the apo-protein form of IscA from Escherichia coli to a resolution of 2.3A. The crystals belong to the space group P3(2)21 and have unit cell dimensions a=b=66.104 A, c=150.167 A (alpha=beta=90 degrees, gamma=120 degrees ). The structure was solved using single-wavelength anomalous dispersion (SAD) phasing of a selenomethionyl derivative, and the IscA model was refined to R=21.4% (Rfree=25.4%). IscA exists as an (alpha1alpha2)2 homotetramer with the (alpha1alpha2) dimer comprising the asymmetric unit. Cys35, implicated in Fe-S cluster assembly, is located in a central cavity formed at the tetramer interface with the gamma-sulfur atoms of residues from the alpha1 and alpha2' monomers (and alpha1'alpha2) positioned close to one another (approximately equal 7 A). C-terminal residues 99-107 are disordered, and the exact positions of Cys99 and Cys101 could not be determined. However, computer modeling of C-terminal residues in the tetramer suggests that Cys99 and Cys101 in the alpha1 monomer and those of the alpha1' monomer (or alpha2 and alpha2') are positioned sufficiently close to coordinate [2Fe-2S] clusters between the two dimers, whereas this is not possible within the (alpha1alpha2) or (alpha1'alpha2') dimer. This symmetrical arrangement allows for binding of two [2Fe-2S] clusters on opposite sides of the tetramer. Modeling further reveals that Cys101 is positioned sufficiently close to Cys35 to allow Cys35 to participate in cluster assembly, formation, or transfer.
- Department of Physiology and Biophysics, University of California, Irvine, CA 92697, USA. jvickery@uci.edu
Organizational Affiliation: