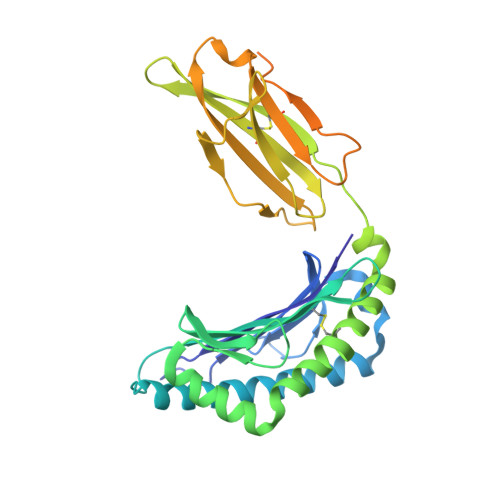
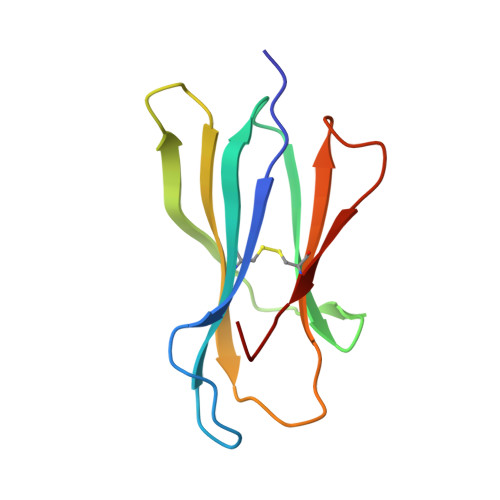
Determination of structural principles underlying three different modes of lymphocytic choriomeningitis virus escape from CTL recognition.
Velloso, L.M., Michaelsson, J., Ljunggren, H.G., Schneider, G., Achour, A.(2004) J Immunol 172: 5504-5511
- PubMed: 15100292 Search on PubMed
- DOI: https://doi.org/10.4049/jimmunol.172.9.5504
- Primary Citation Related Structures:
1S7Q, 1S7R, 1S7S, 1S7T, 1S7U, 1S7V, 1S7W, 1S7X - PubMed Abstract:
Lymphocytic choriomeningitis virus infection of H-2(b) mice generates a strong CD8(+) CTL response mainly directed toward three immunodominant epitopes, one of which, gp33, is presented by both H-2D(b) and H-2K(b) MHC class I molecules. This CTL response acts as a selective agent for the emergence of viral escape variants. These variants generate altered peptide ligands (APLs) that, when presented by class I MHC molecules, antagonize CTL recognition and ultimately allow the virus to evade the cellular immune response. The emergence of APLs of the gp33 epitope is particularly advantageous for LCMV, as it allows viral escape in the context of both H-2D(b) and H-2K(b) MHC class I molecules. We have determined crystal structures of three different APLs of gp33 in complex with both H-2D(b) and H-2K(b). Comparison between these APL/MHC structures and those of the index gp33 peptide/MHC reveals the structural basis for three different strategies used by LCMV viral escape mutations: 1) conformational changes in peptide and MHC residues that are potential TCR contacts, 2) impairment of APL binding to the MHC peptide binding cleft, and 3) introduction of subtle changes at the TCR/pMHC interface, such as the removal of a single hydroxyl group.
- Department of Medical Biochemistry and Biophysics, Karolinska Institute, Stockholm, Sweden.
Organizational Affiliation: