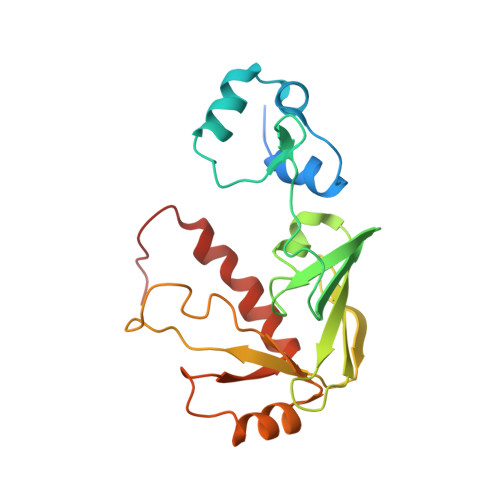
NMR structural studies reveal a novel protein fold for MerB, the organomercurial lyase involved in the bacterial mercury resistance system.
Di Lello, P., Benison, G.C., Valafar, H., Pitts, K.E., Summers, A.O., Legault, P., Omichinski, J.G.(2004) Biochemistry 43: 8322-8332
- PubMed: 15222745 Search on PubMed
- DOI: https://doi.org/10.1021/bi049669z
- Primary Citation Related Structures:
1S6L - PubMed Abstract:
Mercury resistant bacteria have developed a system of two enzymes (MerA and MerB), which allows them to efficiently detoxify both ionic and organomercurial compounds. The organomercurial lyase (MerB) catalyzes the protonolysis of the carbon-mercury bond resulting in the formation of ionic mercury and a reduced hydrocarbon. The ionic mercury [Hg(II)] is subsequently reduced to the less reactive elemental mercury [Hg(0)] by a specific mercuric reductase (MerA). To better understand MerB's unique enzymatic activity, we used nuclear magnetic resonance (NMR) spectroscopy to determine the structure of the free enzyme. MerB is characterized by a novel protein fold consisting of three noninteracting antiparallel beta-sheets surrounded by six alpha-helices. By comparing the NMR data of free MerB and the MerB/Hg/DTT complex, we identified a set of residues that likely define a Hg/DTT binding site. These residues cluster around two cysteines (C(96) and C(159)) that are crucial to MerB's catalytic activity. A detailed analysis of the structure revealed the presence of an extensive hydrophobic groove adjacent to this Hg/DTT binding site. This extensive hydrophobic groove has the potential to interact with the hydrocarbon moiety of a wide variety of substrates and may explain the broad substrate specificity of MerB.
- Department of Biochemistry & Molecular Biology, University of Georgia, Athens, Georgia 30602, USA.
Organizational Affiliation: