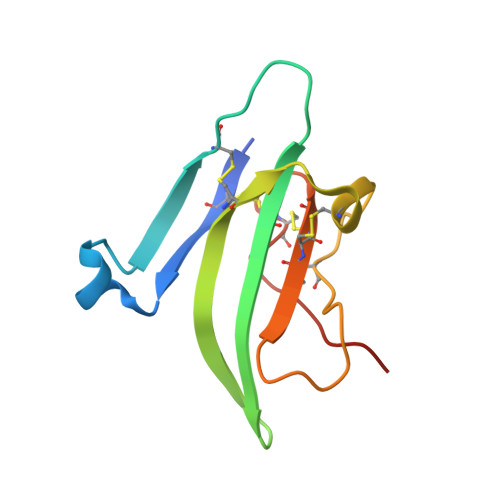
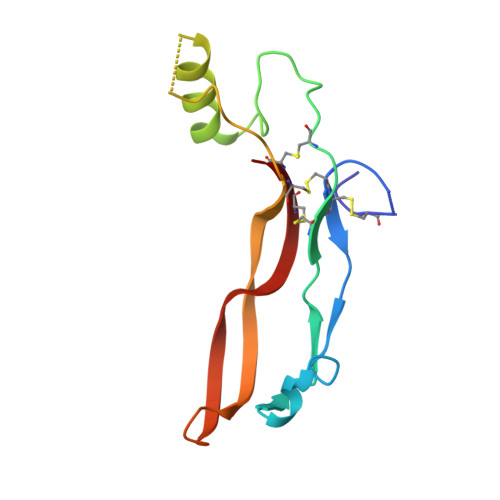
A Flexible Activin Explains the Membrane-Dependent Cooperative Assembly of TGF-beta Family Receptors.
Greenwald, J., Vega, M.E., Allendorph, G.P., Fischer, W.H., Vale, W., Choe, S.(2004) Mol Cell 15: 485-489
- PubMed: 15304227 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2004.07.011
- Primary Citation Related Structures:
1S4Y - PubMed Abstract:
A new crystal structure of activin in complex with the extracellular domain of its type II receptor (ActRIIb-ECD) shows that the ligand exhibits an unexpected flexibility. The motion in the activin dimer disrupts its type I receptor interface, which may account for the disparity in its affinity for type I versus type II receptors. We have measured the affinities of activin and its antagonist inhibin for ActRIIb-ECD and found that the affinity of the 2-fold symmetric homodimer activin for ActRIIb-ECD depends on the availability of two spatially coupled ActRIIb-ECD molecules, whereas the affinity of the heterodimer inhibin does not. Our results indicate that activin's affinity for its two receptor types is greatly influenced by their membrane-restricted setting. We propose that activin affinity is modulated by the ligand flexibility and that cooperativity is achieved by binding to two ActRII chains that immobilize activin in a type I binding-competent orientation.
- Structural Biology Laboratory, The Salk Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: