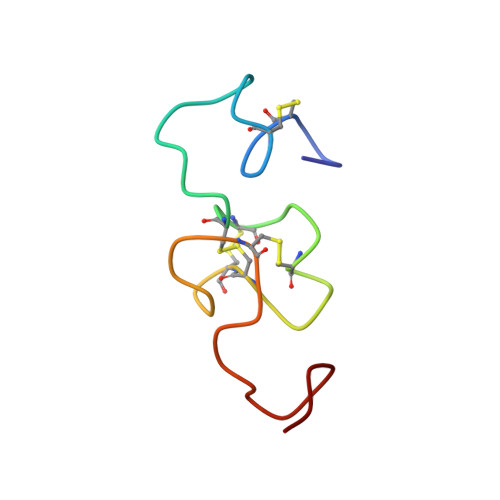
The solution structure of the N-terminal domain of human vitronectin: proximal sites that regulate fibrinolysis and cell migration
Mayasundari, A., Whittemore, N.A., Serpersu, E.H., Peterson, C.B.(2004) J Biological Chem 279: 29359-29366
- PubMed: 15123712 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M401279200
- Primary Citation Related Structures:
1S4G - PubMed Abstract:
The three-dimensional structure of an N-terminal fragment comprising the first 51 amino acids from human plasma vitronectin, the somatomedin B (SMB) domain, has been determined by two-dimensional NMR approaches. An average structure was calculated, representing the overall fold from a set of 20 minimized structures. The core residues (18-41) overlay with a root mean square deviation of 2.29 +/- 0.62 A. The N- and C-terminal segments exhibit higher root mean square deviations, reflecting more flexibility in solution and/or fewer long-range NOEs for these regions. Residues 26-30 form a unique single-turn alpha-helix, the locus where plasminogen activator inhibitor type-1 (PAI-1) is bound. This structure of this helix is highly homologous with that of a recombinant SMB domain solved in a co-crystal with PAI-1 (Zhou, A., Huntington, J. A., Pannu, N. S., Carrell, R. W., and Read, R. J. (2003) Nat. Struct. Biol. 10, 541-544), although the remainder of the structure differs. Significantly, the pattern of disulfide cross-links observed in this material isolated from human plasma is altogether different from the disulfides proposed for recombinant forms. The NMR structure reveals the relative orientation of binding sites for cell surface receptors, including an integrin-binding site at residues 45-47, which was disordered and did not diffract in the co-crystal, and a site for the urokinase receptor, which overlaps with the PAI-1-binding site.
- Department of Biochemistry and Cellular and Molecular Biology and the Center of Excellence in Structural Biology, University of Tennessee, Knoxville, Tennessee 37996, USA.
Organizational Affiliation: