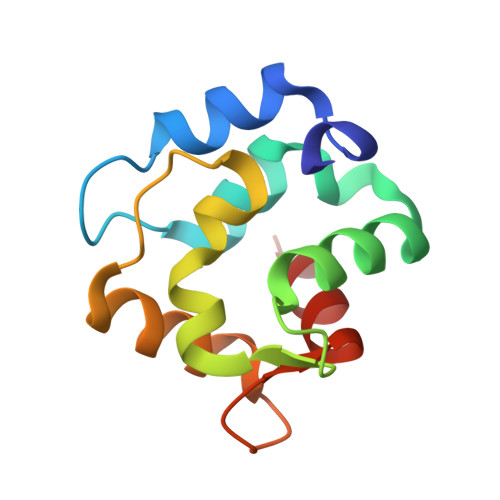
Crystal Structure of a High-Affinity Variant of Rat alpha-Parvalbumin.
Lee, Y.H., Tanner, J.J., Larson, J.D., Henzl, M.T.(2004) Biochemistry 43: 10008-10017
- PubMed: 15287728 Search on PubMed
- DOI: https://doi.org/10.1021/bi0492915
- Primary Citation Related Structures:
1S3P - PubMed Abstract:
In model peptide systems, Ca2+ affinity is maximized in EF-hand motifs containing four carboxylates positioned on the +x and -x and +z and -z axes; introduction of a fifth carboxylate ligand reduces the affinity. However, in rat beta-parvalbumin, replacement of Ser-55 with aspartate heightens divalent ion affinity [Henzl, M. T., et al. (1996) Biochemistry 35, 5856-5869]. The corresponding alpha-parvalbumin variant (S55D/E59D) likewise exhibits elevated affinity [Henzl, M. T., et al. (2003) Anal. Biochem. 319, 216-233]. To determine whether these mutations produce a variation on the archetypal EF-hand coordination scheme, we have obtained high-resolution X-ray crystallographic data for alpha S55D/E59D. As anticipated, the aspartyl carboxylate replaces the serine hydroxyl at the +z coordination position. Interestingly, the Asp-59 carboxylate abandons the role it plays as an outer sphere ligand in wild-type rat beta, rotating away from the Ca2+ and, instead, forming a hydrogen bond with the amide of Glu-62. Superficially, the coordination sphere in the CD site of alpha S55D/E59D resembles that in the EF site. However, the orientation of the Asp-59 side chain is predicted to stabilize the D-helix, which may contribute to the heightened divalent ion affinity. DSC data indicate that the alpha S55D/E59D variant retains the capacity to bind 1 equiv of Na+. Consistent with this finding, when binding measurements are conducted in K(+)-containing buffer, divalent ion affinity is markedly higher. In 0.15 M KCl and 0.025 M Hepes-KOH (pH 7.4) at 5 degrees C, the macroscopic Ca2+ binding constants are 1.8 x 10(10) and 2.0 x 10(9) M(-1). The corresponding Mg2+ binding constants are 2.7 x 10(6) and 1.2 x 10(5) M(-1).
- Department of Chemistry, University of Missouri, Columbia, Missouri 65211, USA.
Organizational Affiliation: