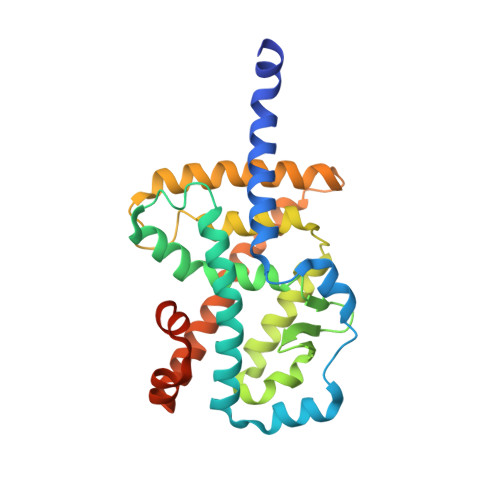
Crystal structure of the human RORalpha Ligand binding domain in complex with cholesterol sulfate at 2.2 A
Kallen, J., Schlaeppi, J.M., Bitsch, F., Delhon, I., Fournier, B.(2004) J Biological Chem 279: 14033-14038
- PubMed: 14722075 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M400302200
- Primary Citation Related Structures:
1S0X - PubMed Abstract:
The retinoic acid-related orphan receptor alpha (RORalpha) is an orphan member of the subfamily 1 of nuclear hormone receptors. Our recent structural and functional studies have led to the hypothesis that cholesterol or a cholesterol derivative is the natural ligand of RORalpha. We have now solved the x-ray crystal structure of the ligand binding domain of RORalpha in complex with cholesterol-3-O-sulfate following a ligand exchange experiment. In contrast to the 3-hydroxyl of cholesterol, the 3-O-sulfate group makes additional direct hydrogen bonds with three residues of the RORalpha ligand binding domain, namely NH-Gln(289), NH-Tyr(290), and NH1-Arg(370). When compared with the complex with cholesterol, seven well ordered water molecules have been displaced, and the ligand is slightly shifted toward the hydrophilic part of the ligand binding pocket, which is ideally suited for interactions with a sulfate group. These additional ligand-protein interactions result in an increased affinity of cholesterol sulfate when compared with cholesterol, as shown by mass spectrometry analysis done under native conditions and differential scanning calorimetry. Moreover, mutational studies show that the higher binding affinity of cholesterol sulfate translates into an increased transcriptional activity of RORalpha. Our findings suggest that cholesterol sulfate could play a crucial role in the regulation of RORalpha in vivo.
- Discovery Technologies, Protein Structure Unit, Basel, Switzerland. joerg.kallen@pharma.novartis.com
Organizational Affiliation: