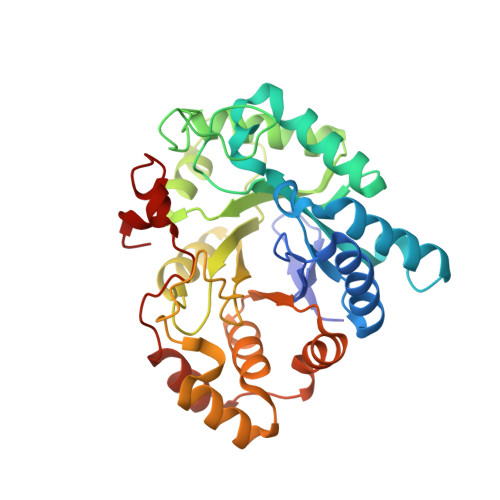
Crystal structure of human prostaglandin F synthase (AKR1C3).
Komoto, J., Yamada, T., Watanabe, K., Takusagawa, F.(2004) Biochemistry 43: 2188-2198
- PubMed: 14979715 Search on PubMed
- DOI: https://doi.org/10.1021/bi036046x
- Primary Citation Related Structures:
1RY0, 1RY8 - PubMed Abstract:
Prostaglandin H(2) (PGH(2)) formed from arachidonic acid is an unstable intermediate and is efficiently converted into more stable arachidonate metabolites (PGD(2), PGE(2), and PGF(2)) by the action of three groups of enzymes. Prostaglandin F synthase (PGFS) was first purified from bovine lung and catalyzes the formation of 9 alpha,11 beta-PGF(2) from PGD(2) and PGF(2)(alpha) from PGH(2) in the presence of NADPH. Human PGFS is 3 alpha-hydroxysteroid dehydrogenase (3 alpha-HSD) type II and has PGFS activity and 3 alpha-HSD activity. Human lung PGFS has been crystallized with the cofactor NADP(+) and the substrate PGD(2), and with the cofactor NADPH and the inhibitor rutin. These complex structures have been determined at 1.69 A resolution. PGFS has an (alpha/beta)(8) barrel structure. The cofactor and substrate or inhibitor bind in a cavity at the C-terminal end of the barrel. The cofactor binds deeply in the cavity and has extensive interactions with PGFS through hydrogen bonds, whereas the substrate (PGD(2)) is located above the bound cofactor and has little interaction with PGFS. Despite being largely structurally different from PGD(2), rutin is located at the same site of PGD(2), and its catechol and rhamnose moieties are involved in hydrogen bonds with PGFS. The catalytic site of PGFS contains the conserved Y55 and H117 residues. The carbonyl O(11) of PGD(2) and the hydroxyl O(13) of rutin are involved in hydrogen bonds with Y55 and H117. The cyclopentane ring of PGD(2) and the phenyl ring of rutin face the re-side of the nicotinamide ring of the cofactor. On the basis of the catalytic geometry, a direct hydride transfer from NADPH to PGD(2) would be a reasonable catalytic mechanism. The hydride transfer is facilitated by protonation of carbonyl O(11) of PGD(2) from either H117 (at low pH) or Y55 (at high pH). Since the substrate binding cavity of PGFS is relatively large in comparison with those of AKR1C1 and AKR1C2, PGFS (AKR1C3) could catalyze the reduction and/or oxidation reactions of various compounds over a relatively wide pH range.
- Department of Molecular Biosciences, University of Kansas, 1200 Sunnyside Avenue, Lawrence, Kansas 66045-7534, USA.
Organizational Affiliation: