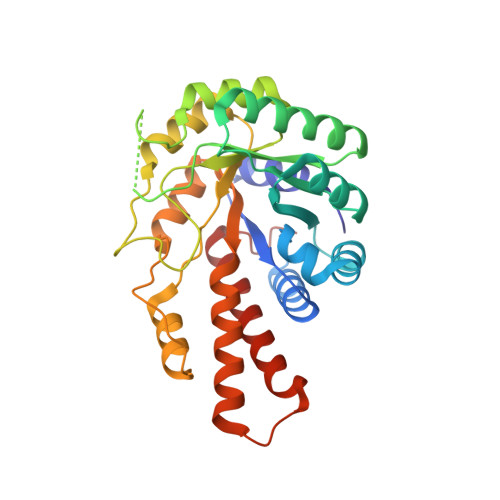
Induced Fit Movements and Metal Cofactor Selectivity of Class II Aldolases: STRUCTURE OF THERMUS AQUATICUS FRUCTOSE-1,6-BISPHOSPHATE ALDOLASE.
Izard, T., Sygusch, J.(2004) J Biological Chem 279: 11825-11833
- PubMed: 14699122 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M311375200
- Primary Citation Related Structures:
1RV8, 1RVG - PubMed Abstract:
Fructose-1,6-bisphosphate (FBP) aldolase is an essential glycolytic enzyme that reversibly cleaves its ketohexose substrate into triose phosphates. Here we report the crystal structure of a metallo-dependent or class II FBP aldolase from an extreme thermophile, Thermus aquaticus (Taq). The quaternary structure reveals a tetramer composed of two dimers related by a 2-fold axis. Taq FBP aldolase subunits exhibit two distinct conformational states corresponding to loop regions that are in either open or closed position with respect to the active site. Loop closure remodels the disposition of chelating active site histidine residues. In subunits corresponding to the open conformation, the metal cofactor, Co(2+), is sequestered in the active site, whereas for subunits in the closed conformation, the metal cation exchanges between two mutually exclusive binding loci, corresponding to a site at the active site surface and an interior site vicinal to the metal-binding site in the open conformation. Cofactor site exchange is mediated by rotations of the chelating histidine side chains that are coupled to the prior conformational change of loop closure. Sulfate anions are consistent with the location of the phosphate-binding sites of the FBP substrate and determine not only the previously unknown second phosphate-binding site but also provide a mechanism that regulates loop closure during catalysis. Modeling of FBP substrate into the active site is consistent with binding by the acyclic keto form, a minor solution species, and with the metal cofactor mediating keto bond polarization. The Taq FBP aldolase structure suggests a structural basis for different metal cofactor specificity than in Escherichia coli FBP aldolase structures, and we discuss its potential role during catalysis. Comparison with the E. coli structure also indicates a structural basis for thermostability by Taq FBP aldolase.
- Department of Hematology-Oncology, St. Jude Children's Research Hospital, Memphis, Tennessee 38111, USA.
Organizational Affiliation: