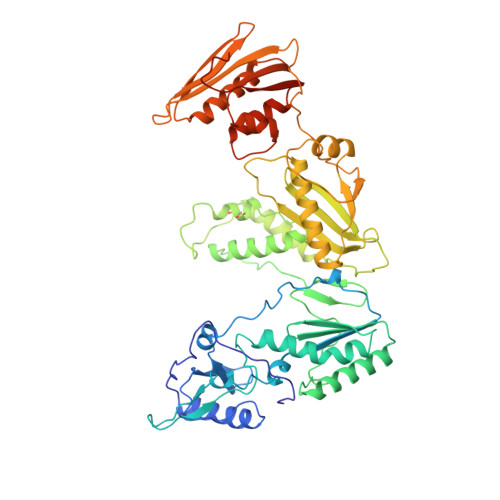
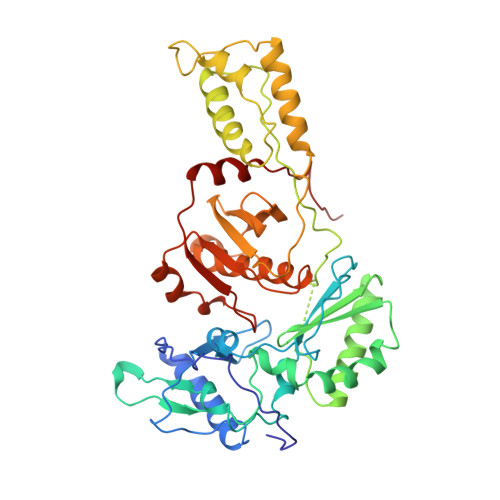
Mechanism of inhibition of HIV-1 reverse transcriptase by non-nucleoside inhibitors.
Esnouf, R., Ren, J., Ross, C., Jones, Y., Stammers, D., Stuart, D.(1995) Nat Struct Biol 2: 303-308
- PubMed: 7540935 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0495-303
- Primary Citation Related Structures:
1RTJ - PubMed Abstract:
The structure of unliganded HIV-1 reverse transcriptase has been determined at 2.35 A resolution and refined to an R-factor of 0.219 (for all data) with good stereochemistry. The unliganded structure was produced by soaking out a weak binding non-nucleoside inhibitor, HEPT, from pregrown crystals. Comparison with the structures of four different RT and non-nucleoside inhibitor complexes reveals that only minor domain rearrangements occur, but there is a significant repositioning of a three-stranded beta-sheet in the p66 subunit (containing the catalytic aspartic acid residues 110, 185 and 186) with respect to the rest of the polymerase site. This suggests that NNIs inhibit RT by locking the polymerase active site in an inactive conformation, reminiscent of the conformation observed in the inactive p51 subunit.
- Laboratory of Molecular Biophysics, Oxford, UK.
Organizational Affiliation: