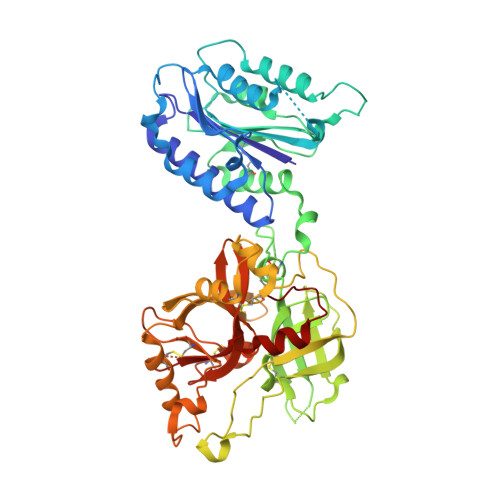
Structural analysis of engineered Bb fragment of complement factor B: insights into the activation mechanism of the alternative pathway C3-convertase.
Ponnuraj, K., Xu, Y., Macon, K., Moore, D., Volanakis, J.E., Narayana, S.V.(2004) Mol Cell 14: 17-28
- PubMed: 15068800 Search on PubMed
- DOI: https://doi.org/10.1016/s1097-2765(04)00160-1
- Primary Citation Related Structures:
1RRK, 1RS0, 1RTK - PubMed Abstract:
The C-terminal fragment, Bb, of factor B combines with C3b to form the pivotal C3-convertase, C3bBb, of alternative complement pathway. Bb consists of a von Willebrand factor type A (vWFA) domain that is structurally similar to the I domains of integrins and a serine protease (SP) domain that is in inactive conformation. The structure of the C3bBb complex would be important in deciphering the activation mechanism of the SP domain. However, C3bBb is labile and not amenable to X-ray diffraction studies. We engineered a disulfide bond in the vWFA domain of Bb homologous to that shown to lock I domains in active conformation. The crystal structures of Bb(C428-C435) and its inhibitor complexes reveal that the adoption of the "active" conformation by the vWFA domain is not sufficient to activate the C3-convertase catalytic apparatus and also provide insights into the possible mode of C3-convertase activation.
- Center for Biophysical Sciences and Engineering, School of Optometry, University of Alabama at Birmingham, Birmingham, AL 35294, USA.
Organizational Affiliation: