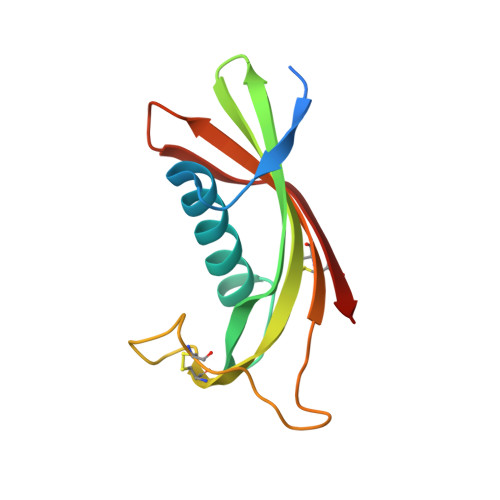
Crystal structure of human cystatin D, a cysteine peptidase inhibitor with restricted inhibition profile.
Alvarez-Fernandez, M., Liang, Y.H., Abrahamson, M., Su, X.D.(2005) J Biological Chem 280: 18221-18228
- PubMed: 15728581 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M411914200
- Primary Citation Related Structures:
1RN7, 1ROA - PubMed Abstract:
Cystatins are natural inhibitors of papain-like (family C1) and legumain-related (family C13) cysteine peptidases. Cystatin D is a type 2 cystatin, a secreted inhibitor found in human saliva and tear fluid. Compared with its homologues, cystatin D presents an unusual inhibition profile with a preferential inhibition cathepsin S > cathepsin H > cathepsin L and no inhibition of cathepsin B or pig legumain. To elucidate the structural reasons for this specificity, we have crystallized recombinant human Arg(26)-cystatin D and solved its structures at room temperature and at cryo conditions to 2.5- and 1.8-A resolution, respectively. Human cystatin D presents the typical cystatin fold, with a five-stranded anti-parallel beta-sheet wrapped around a five-turn alpha-helix. The structures reveal differences in the peptidase-interacting regions when compared with other cystatins, providing plausible explanations for the restricted inhibitory specificity of cystatin D for some papain-like peptidases and its lack of reactivity toward legumain-related enzymes.
- Department of Clinical Chemistry, Institute of Laboratory Medicine, Lund University, SE-221 85 Lund, Sweden.
Organizational Affiliation: