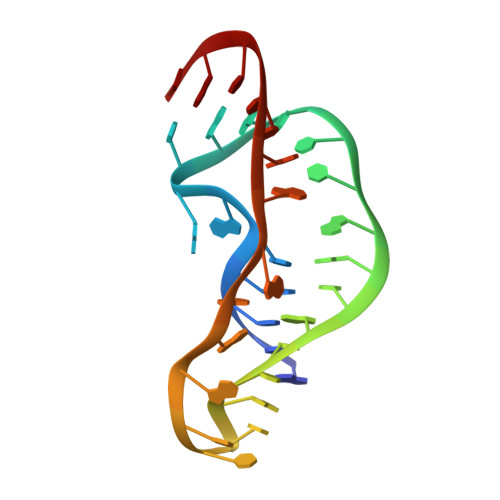
The structure of an RNA pseudoknot that causes efficient frameshifting in mouse mammary tumor virus.
Shen, L.X., Tinoco Jr., I.(1995) J Mol Biology 247: 963-978
- PubMed: 7723043 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1995.0193
- Primary Citation Related Structures:
1RNK - PubMed Abstract:
The structure of a 34-nucleotide RNA pseudoknot that causes efficient -1 frameshifting in the messenger RNA of mouse mammary tumor virus has been investigated by NMR. Spectral assignment of the pseudoknot was facilitated by comparative NMR studies on the pseudoknot and on two smaller hairpin RNAs, and by using selective 13C labeling and 13C-edited NMR techniques. The three-dimensional structure of the pseudoknot has been determined. The frameshifter pseudoknot possesses structural features not observed in previously reported model pseudoknots. It has a compact structure with a pronounced bend at the junction of its G.C-rich stems. A single adenylate residue is intercalated between the two stems so that direct coaxial staking of the stems is not possible. The lack of an opposing nucleotide for the stacked, intervening adenylate creates a hinge in the pseudoknot. Most of the loop nucleotides are restrained by base staking interactions which keep the loops from adopting extended conformations. The sterically constrained loops direct the bending of the pseudoknot at the stem-stem junction. The roles of the intercalated adenylate and loop lengths in causing bending can explain their requirement for efficient frameshifting. Our NMR data also indicate that there are internal dynamics associated with the pseudoknot. The unique, compact structure and conformational flexibility of the pseudoknot may be required for recognition and favourable interaction with the translating ribosome, or with translation factors associated with the ribosome.
- Department of Chemistry, University of California, Berkeley, USA.
Organizational Affiliation: