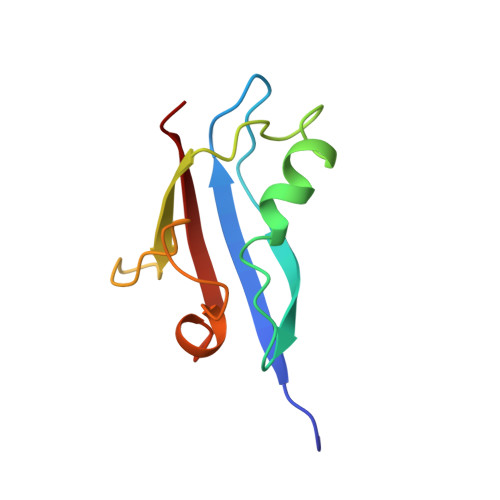
Structure determination of the Ras-binding domain of the Ral-specific guanine nucleotide exchange factor Rlf.
Esser, D., Bauer, B., Wolthuis, R.M., Wittinghofer, A., Cool, R.H., Bayer, P.(1998) Biochemistry 37: 13453-13462
- PubMed: 9753431 Search on PubMed
- DOI: https://doi.org/10.1021/bi9811664
- Primary Citation Related Structures:
1RLF - PubMed Abstract:
Ral-specific guanine nucleotide exchange factors RalGDS, Rgl, and Rlf have been suggested to function as intermediates between Ras and Ral pathways by being able to bind Ras proteins through their C-terminal Ras-binding domains (RBD). The RBDs of RalGDS and of the Ser/Thr kinase c-Raf-1 have been shown to have the same tertiary structure. In contrast to the RBDs of Raf and RalGDS, which bind either Ras or Rap with high affinity, Rlf-RBD has a similar affinity for both GTP-binding proteins. To be able to compare these RBDs on a structural level, we have solved the three-dimensional structure of Rlf-RBD by NMR spectroscopy. The overall tertiary structure of Rlf-RBD shows the betabetaalphabetabetaalphabeta-fold of the ubiquitin superfamily and is very similar to that of RalGDS-RBD. The binding interface of Rlf-RBD to Ras was mapped using chemical shift analysis and indicated a binding mode similar to that in the case of Rap.Raf-RBD. However, comparison of the putatively interacting regions revealed structural differences which are proposed to be responsible for the different substrate affinities of Rlf-, RalGDS-, and Raf-RBD.
- Max-Planck-Institut für molekulare Physiologie, Abteilung Strukturelle Biologie, Abteilung Physikalische Biochemie, Dortmund, Germany.
Organizational Affiliation: