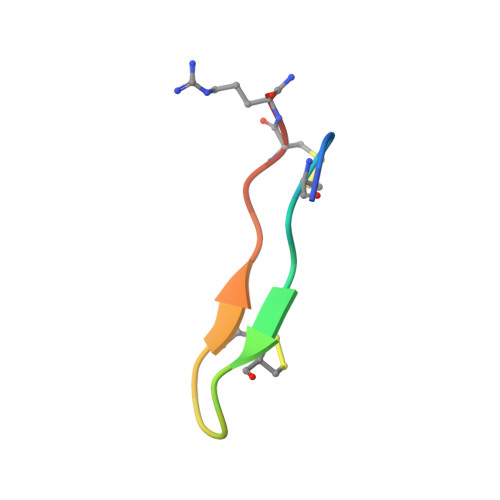
Structure-activity relationships for the beta-hairpin cationic antimicrobial peptide polyphemusin I.
Powers, J.P.S., Rozek, A., Hancock, R.E.W.(2004) Biochim Biophys Acta 1698: 239-250
- PubMed: 15134657
- DOI: https://doi.org/10.1016/j.bbapap.2003.12.009
- Primary Citation of Related Structures:
1RKK - PubMed Abstract:
The solution structure of polyphemusin I was determined using (1)H-NMR spectroscopy. Polyphemusin I was found to be an amphipathic, beta-hairpin connected by a type I' beta-turn. The 17 low-energy structures aligned very well over the beta-sheet region while both termini were poorly defined due in part to a hinge-like region centred in the molecule about arginine residues 6 and 16. Conversely, a linear analogue, PM1-S, with all cysteines simultaneously replaced with serine was found to be dynamic in nature, and a lack of medium and long-range NOEs indicated that this molecule displayed no favoured conformation. Circular dichroism (CD) spectroscopy confirmed that in solution, 50% trifluoroethanol (TFE) and in the presence of liposomes, PM1-S remained unstructured. The antimicrobial activity of PM1-S was found to be 4- to 16-fold less than that of polyphemusin I and corresponded with a 4-fold reduction in bacterial membrane depolarization. Both peptides were able to associate with lipid bilayers in a similar fashion; however, PM1-S was completely unable to translocate model membranes while polyphemusin I retained this activity. It was concluded that the disulfide-constrained, beta-sheet structure of polyphemusin I is required for maximum antimicrobial activity. Disruption of this structure results in reduced antimicrobial activity and completely abolishes membrane translocation indicating that the linear PM1-S acts through a different antimicrobial mechanism.
- Department of Microbiology and Immunology, University of British Columbia, #300-6174 University Boulevard, Vancouver, British Columbia, V6T 1Z3, Canada.
Organizational Affiliation: