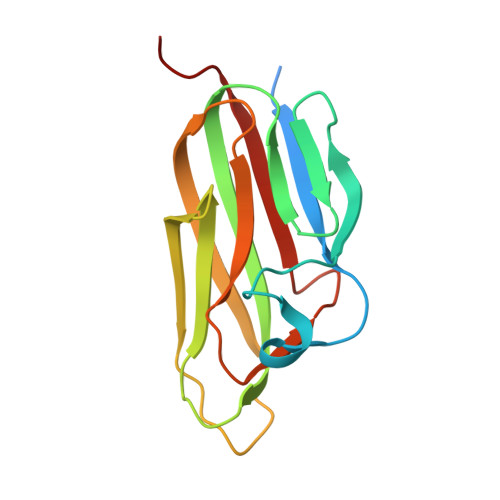
The crystal structures of EDA-A1 and EDA-A2: splice variants with distinct receptor specificity.
Hymowitz, S.G., Compaan, D.M., Yan, M., Wallweber, H.J., Dixit, V.M., Starovasnik, M.A., de Vos, A.M.(2003) Structure 11: 1513-1520
- PubMed: 14656435 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2003.11.009
- Primary Citation Related Structures:
1RJ7, 1RJ8 - PubMed Abstract:
EDA is a tumor necrosis factor family member involved in ectodermal development. Splice variants EDA-A1 and EDA-A2 differ only by the presence of Glu 308 and Val 309 in the expected receptor binding region of EDA-A1 but not EDA-A2. This two amino acid difference functions as a switch controlling receptor specificity. EDA-A1 binds only to EDAR, while EDA-A2 is specific for XEDAR. In order to understand the structural basis of this switch, we determined the X-ray crystal structures of the TNF domain of both EDA-A1 and EDA-A2 at 2.3 A and 2.2 A, respectively. While the backbone conformation around the splice difference is similar in both isoforms, the conformation of the following loop, the surface charge, and the shape of the expected receptor binding site differ significantly.
- Department of Protein Engineering, Genentech, Inc., 1 DNA Way, South San Francisco, CA 94080, USA. hymowitz@gene.com
Organizational Affiliation: