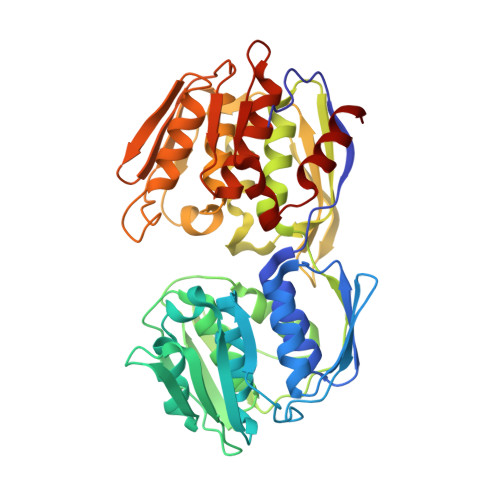
Structural studies of Streptococcus pneumoniae EPSP synthase in unliganded state, tetrahedral intermediate-bound state and S3P-GLP-bound state.
Park, H., Hilsenbeck, J.L., Kim, H.J., Shuttleworth, W.A., Park, Y.H., Evans, J.N., Kang, C.(2004) Mol Microbiol 51: 963-971
- PubMed: 14763973 Search on PubMed
- DOI: https://doi.org/10.1046/j.1365-2958.2003.03885.x
- Primary Citation Related Structures:
1RF4, 1RF5, 1RF6 - PubMed Abstract:
The shikimate pathway synthesizes aromatic amino acids and other essential metabolites that are necessary for bacteria, plants and fungi to survive. This pathway is not present in vertebrates and therefore represents an attractive target for antibacterial agents. We have successfully crystallized and solved the structure of unliganded, inhibitor-liganded and tetrahedral intermediate (TI)-liganded forms of Streptococcus pneumoniae EPSP synthase. The overall topology of the S. pneumoniae EPSP synthase is similar to that of the Escherichia coli EPSP synthase. In addition, the majority of residues responsible for ligand binding were conserved between the two proteins. TI-liganded structure provides absolute configuration of the C-2 atom from the F-PEP moiety of the enzyme-bound intermediate and also defines key residues responsible for the enzyme reaction. Comparison of the unliganded state and substrate-bound state of the enzyme provides insights into the structural mechanisms involved in dynamic events of ligand binding, domain movement and closure. This structural study of the pathogenic bacteria S. pneumoniae EPSP synthase with inhibitor and TI will provide invaluable information for the design of new-generation antibiotics.
- School of Molecular Biosciences, Washington State University, Pullman 99164, USA.
Organizational Affiliation: