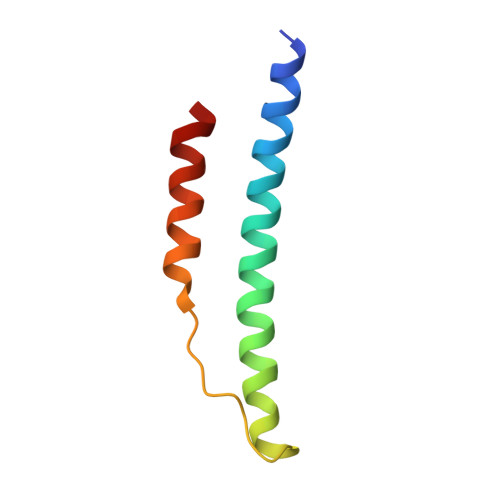
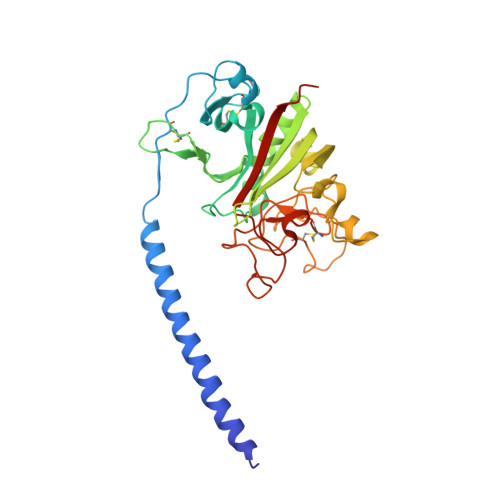
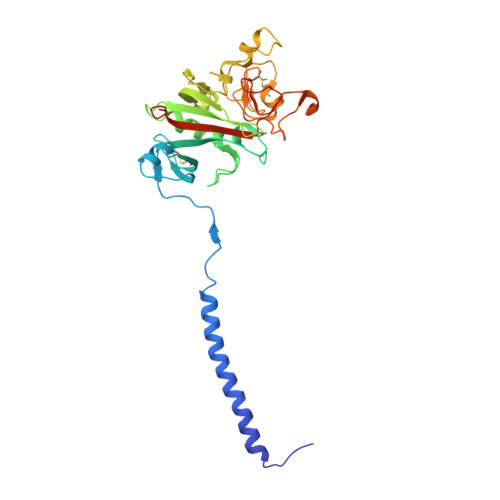
Calcium-Binding Site beta2, Adjacent to the "b" Polymerization Site, Modulates Lateral Aggregation of Protofibrils during Fibrin Polymerization.
Kostelansky, M.S., Lounes, K.C., Ping, L.F., Dickerson, S.K., Gorkun, O.V., Lord, S.T.(2004) Biochemistry 43: 2475-2483
- PubMed: 14992585
- DOI: https://doi.org/10.1021/bi0359978
- Primary Citation of Related Structures:
1RF0, 1RF1 - PubMed Abstract:
Structural analysis of recombinant fibrinogen fragment D revealed that the calcium-binding site (beta2-site) composed of residues BbetaAsp261, BbetaAsp398, BbetaGly263, and gammaGlu132 is modulated by the "B:b" interaction. To determine the beta2-site's role in polymerization, we engineered variant fibrinogen gammaE132A in which calcium binding to the beta2-site was disrupted by replacing glutamic acid at gamma132 with alanine. We compared polymerization of gammaE132A to normal fibrinogen as a function of calcium concentration. Polymerization of gammaE132A at concentrations of calcium
- Department of Chemistry, University of North Carolina, Chapel Hill, North Carolina 27599-7525, USA.
Organizational Affiliation: