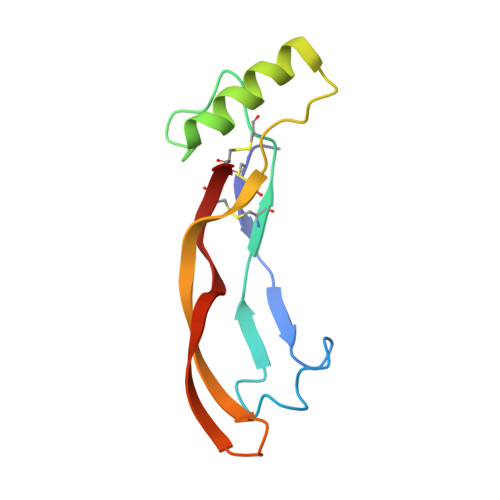
Molecular recognition of BMP-2 and BMP receptor IA.
Keller, S., Nickel, J., Zhang, J.L., Sebald, W., Mueller, T.D.(2004) Nat Struct Mol Biol 11: 481-488
- PubMed: 15064755 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb756
- Primary Citation Related Structures:
1REU, 1REW - PubMed Abstract:
Bone morphogenetic protein-2 (BMP-2) and other members of the TGF-beta superfamily regulate the development, maintenance and regeneration of tissues and organs. Binding epitopes for these extracellular signaling proteins have been defined, but hot spots specifying binding affinity and specificity have so far not been identified. In this study, mutational and structural analyses show that epitopes of BMP-2 and the BRIA receptor form a new type of protein-protein interface. The main chain atoms of Leu 51 and Asp53 of BMP-2 represent a hot spot of binding to BRIA. The BMP-2 variant L51P was deficient in type I receptor binding only, whereas its overall structure and its binding to type II receptors and modulator proteins, such as noggin, were unchanged. Thus, the L51P substitution converts BMP-2 into a receptor-inactive inhibitor of noggin. These results are relevant for other proteins of the TGF-beta superfamily and provide useful clues for structure-based drug design.
- Lehrstuhl für Physiologische Chemie II, Theodor-Boveri Institut für Biowissenschaften (Biozentrum), Universität Würzburg, Am Hubland, D-97074 Wuerzburg, Germany.
Organizational Affiliation: