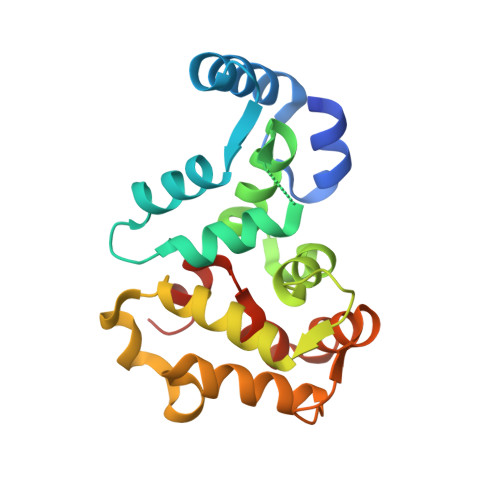
Three-Dimensional Structure of Recoverin, a Calcium Sensor in Vision
Flaherty, K.M., Zozulya, S., Stryer, L., McKay, D.B.(1993) Cell 75: 709-716
- PubMed: 8242744 Search on PubMed
- DOI: https://doi.org/10.1016/0092-8674(93)90491-8
- Primary Citation Related Structures:
1REC - PubMed Abstract:
Recoverin, a recently discovered member of the EF hand superfamily, serves as a calcium sensor in vision. We report here the crystal structure of recombinant unmyristoylated recoverin at 1.9 A resolution. The four EF hands of the protein are arranged in a compact array that contrasts with the dumbbell shape of calmodulin and troponin C. A calcium ion is bound to EF hand 3, while EF hand 2 can bind samarium but not calcium in this crystal form. The other two EF hands have novel structural features that prevent or impair calcium binding. A concave hydrophobic surface formed by EF hands 1 and 2 may participate in the read out of calcium signals by recoverin and its homologs.
- Beckman Laboratories for Structural Biology, Department of Cell Biology, Stanford University School of Medicine, California 94305.
Organizational Affiliation: