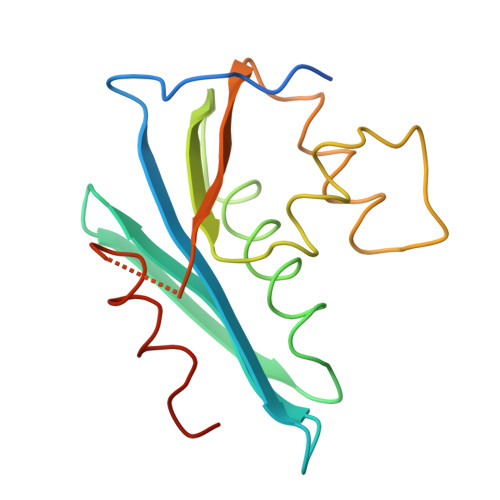
Crystallographic analyses of an active HIV-1 ribonuclease H domain show structural features that distinguish it from the inactive form.
Chattopadhyay, D., Finzel, B.C., Munson, S.H., Evans, D.B., Sharma, S.K., Strakalaitus, N.A., Brunner, D.P., Eckenrode, F.M., Dauter, Z., Betzel, C., Einspahr, H.M.(1993) Acta Crystallogr D Biol Crystallogr 49: 423-427
- PubMed: 15299518 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444993002409
- Primary Citation Related Structures:
1RDH - PubMed Abstract:
. An active recombinant preparation of the carboxy-terminal ribonuclease H (RNase H) domain of HIV-I reverse transcriptase has produced crystals of several different forms, including a trigonal prism form (P3(1); a = b = 52.03, c = 113.9 A with two molecules per asymmetric unit) and a hexagonal tablet form (P6(2)22 or P6(4)22; a = b = 93.5, c = 74.1 A with one molecule per asymmetric unit). The former appears to be isomorphous with crystals of a similar, but inactive, version of the enzyme that was used for a prior crystal structure determination [Davies, Hostomska, Hostomsky, Jordan & Matthews (1991). Science, 252, 88-95]. We have also obtained a structure solution for this crystal form and have refined it with 2.8 A resolution data (R = 0.216). We report here details of our crystallization studies and some initial structural results that verify that the preparation of active HIV-1 RNase H yields a protein that is not just enzymatically, but also structurally, distinguishable from the inactive form. Evidence suggests that region 538-542, which may be involved in the catalytic site and which is disordered in both molecules in the prior structure determination, is ordered in the crystal structure of the active enzyme, although the ordering may include more than one conformation for this loop. It should also be noted that, in the crystal structure of the trigonal form, RNase H monomers associate to form noncrystallographic twofold-symmetric dimers by fusing five-stranded mixed beta sheets into a single ten-stranded dimerwide sheet, an assembly that was not remarked upon by previous investigators.
- The Upjohn Company, Kalamazoo, Michigan 49001, USA.
Organizational Affiliation: