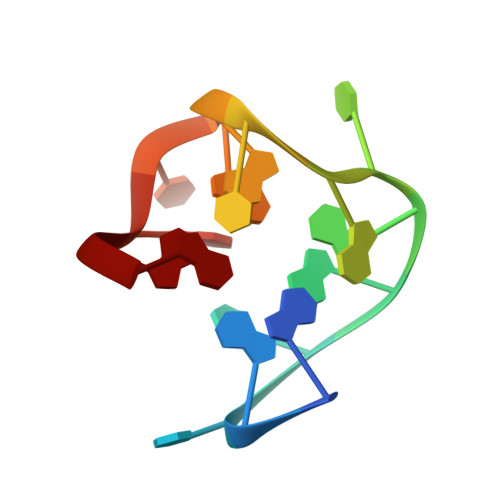
NMR structure of the thrombin-binding DNA aptamer stabilized by Sr2+.
Mao, X., Marky, L.A., Gmeiner, W.H.(2004) J Biomol Struct Dyn 22: 25-33
- PubMed: 15214802 Search on PubMed
- DOI: https://doi.org/10.1080/07391102.2004.10506977
- Primary Citation Related Structures:
1RDE - PubMed Abstract:
The structure of thrombin-binding DNA aptamer complexed with a single Sr2+ ion (Sr2+:TBA complex) has been determined using NMR spectroscopy and restrained molecular dynamics simulations. The quadruplex structure for the Sr2+:TBA complex is similar in topology, but distinct in structure, from that previously reported for the K+:TBA complex. The inter-tetrad distance of the Sr2+:TBA complex is 3.8 angstroms, or 0.7 angstroms larger than in the K+:TBA complex. This substantial difference can be attributed to a different binding site for Sr2+ in the Sr2+:TBA complex than for K+ in the K+:TBA complex. The Sr2+:TBA complex assumes a 1:1 stoichiometry, and it is very likely that the Sr2+ ion simultaneously interacts with the eight O6 atoms of the two G-tetrads. The results indicate that quadruplex DNA structures are highly sensitive to the presence of specific metal ions. The binding of specific metal ions may modulate the biological activity of quadruplex DNA structures in vivo.
- Department of Biochemistry, Wake Forest University School of Medicine, Winston-Salem, NC 27157-1016, USA.
Organizational Affiliation: