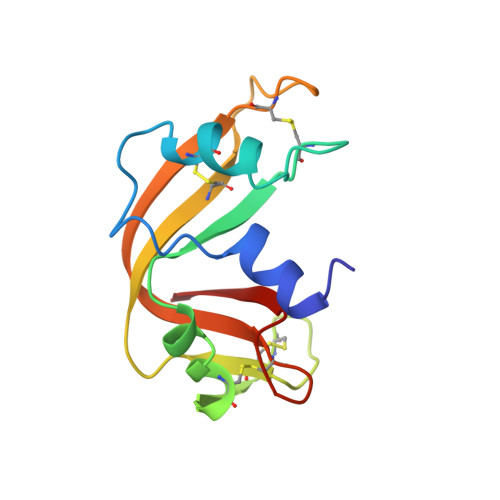
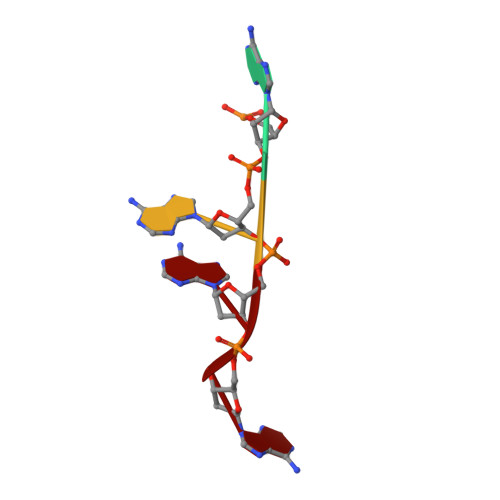
Structure of a ribonuclease B+d(pA)4 complex.
Ko, T.P., Williams, R., McPherson, A.(1996) Acta Crystallogr D Biol Crystallogr 52: 160-164
- PubMed: 15299737 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444995009127
- Primary Citation Related Structures:
1RBJ - PubMed Abstract:
The structure of a tetragonal crystal of bovine pancreatic RNase B complexed with d(pA)(4) was determined by molecular replacement and difference Fourier methods. This crystal belongs to space group P4(1)2(1)2 and has unit-cell dimensions a = b = 44.5, c = 156.5 A. The model consists of the enzyme and a tetranucleotide with fractional occupancies, suggesting multiple modes of oligonucleotide binding. It does not include any polysaccharide residues or solvent molecules. After refinement at 2.7 A, the R value was 0.163 with acceptable stereochemistry. The model illustrates a set of well defined interactions for substrate binding, particularly between the central dinucleotide and the enzyme.
- Department of Biochemistry, University of California, Riverside 92521, USA.
Organizational Affiliation: