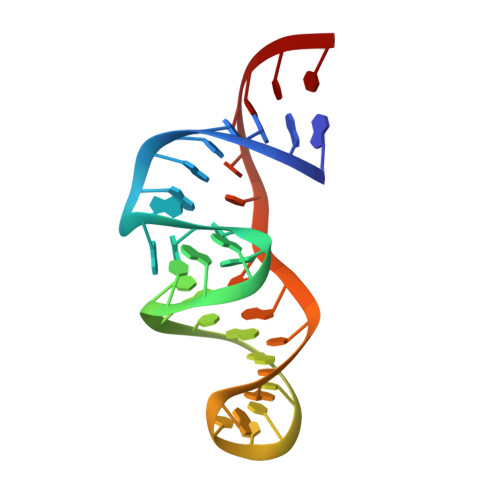Solution Structure of an ATP-Binding RNA Aptamer Reveals a Novel Fold
Dieckmann, T., Suzuki, E., Nakamura, G.K., Feigon, J.(1996) RNA 2: 628-640
- PubMed: 8756406 Search on PubMedSearch on PubMed Central
- Primary Citation Related Structures:
1RAW - PubMed Abstract:
In vitro selection has been used to isolate several RNA aptamers that bind specifically to biological cofactors. A well-characterized example in the ATP-binding RNA aptamer family, which contains a conserved 11-base loop opposite a bulged G and flanked by regions of double-stranded RNA. The nucleotides in the consensus sequence provide a binding pocket for ATP (or AMP), which binds with a Kd in the micromolar range. Here we present the three-dimensional solution structure of a 36-nucleotide ATP-binding RNA aptamer complexed with AMP, determined from NMR-derived distance and dihedral angle restraints. The conserved loop and bulged G form a novel compact, folded structure around the AMP. The backbone tracing of the loop nucleotides can be described by a Greek zeta (zeta). Consecutive loop nucleotides G, A, A form a U-turn at the bottom of the zeta, and interact with the AMP to form a structure similar to a GNRA tetraloop, with AMP standing in for the final A. Two asymmetric G. G base pairs close the stems flanking the internal loop. Mutated aptamers support the existence of the tertiary interactions within the consensus nucleotides and with the AMP found in the calculated structures.
- Department of Chemistry and Biochemistry, University of California, Los Angeles 90095, USA.
Organizational Affiliation: