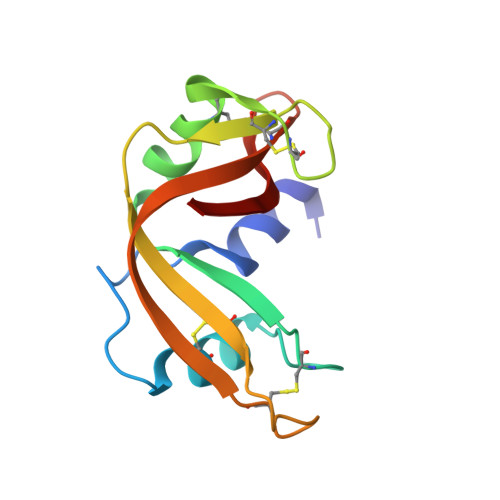
Crystal structure of a fluorescent derivative of RNase A.
Baudet-Nessler, S., Jullien, M., Crosio, M.P., Janin, J.(1993) Biochemistry 32: 8457-8464
- PubMed: 8357795 Search on PubMed
- Primary Citation Related Structures:
1RAR, 1RAS - PubMed Abstract:
The crystal structure of RNase A chemically modified with the fluorescent probe, N-[[(iodoacetyl)-amino]ethyl]-5-naphthylamine-1-sulfonic acid (1,5-IAENS), has been solved and refined to high resolution. It yields information on the mode of binding, the mobility of a probe commonly used in spectroscopic studies, and anion binding sites in RNase A. Trigonal crystals of the fluorescent derivative grown in sodium or cesium chloride and ammonium sulfate, pH 5.1, were nearly isomorphous with those of a semisynthetic RNase [DeMel, et al. (1992) J. Biol. Chem. 267, 247-256]. Refinement starting from semisynthetic RNase led to a model with R = 20% against 1.7-A diffraction data from crystals in ammonium sulfate and another model with R = 17% against 1.9-A data taken in the presence of 3 M NaCl. The second model contains three chloride ions: one is at the active site, and the other two are at molecular interfaces. Otherwise, the two models are very similar. The fluorophore has very little effect on the protein conformation. It is found to be covalently attached to the active site His-12 with the naphthyl group stacked on the imidazole ring of His-119. It remains largely accessible to solvent and in a polar environment on the protein surface, even though the fluorescence emission spectrum is blue shifted as it is in nonpolar solvents.
- Laboratoire de Biologie Structurale, UMR 9920 CNRS-Université Paris-Sud, Orsay, France.
Organizational Affiliation: