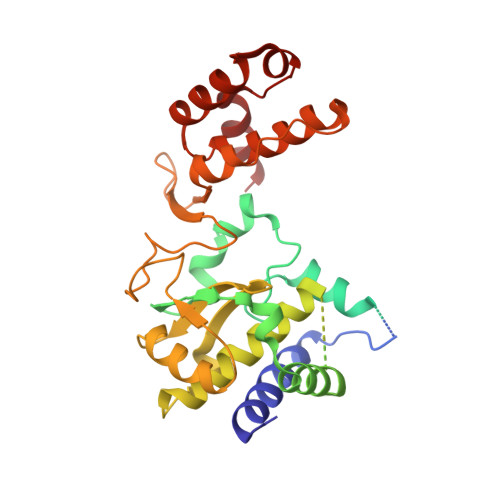
Structural basis for proteolysis-dependent activation of the poliovirus RNA-dependent RNA polymerase.
Thompson, A.A., Peersen, O.B.(2004) EMBO J 23: 3462-3471
- PubMed: 15306852 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7600357
- Primary Citation Related Structures:
1RA6, 1RA7, 1RAJ, 1TQL - PubMed Abstract:
The active RNA-dependent RNA polymerase of poliovirus, 3Dpol, is generated by cleavage of the 3CDpro precursor protein, a protease that has no polymerase activity despite containing the entire polymerase domain. By intentionally disrupting a known and persistent crystal packing interaction, we have crystallized the poliovirus polymerase in a new space group and solved the complete structure of the protein at 2.0 A resolution. It shows that the N-terminus of fully processed 3Dpol is buried in a surface pocket where it makes hydrogen bonds that act to position Asp238 in the active site. Asp238 is an essential residue that selects for the 2' OH group of substrate rNTPs, as shown by a 2.35 A structure of a 3Dpol-GTP complex. Mutational, biochemical, and structural data further demonstrate that 3Dpol activity is exquisitely sensitive to mutations at the N-terminus. This sensitivity is the result of allosteric effects where the structure around the buried N-terminus directly affects the positioning of Asp238 in the active site.
- Program in Cellular and Molecular Biology, Colorado State University, Fort Collins 80523, USA.
Organizational Affiliation: