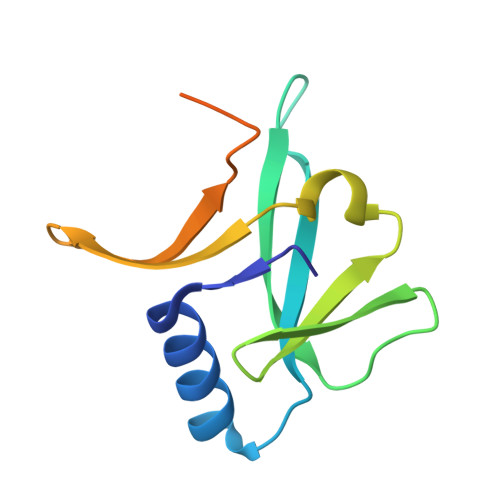
Crystal structure of the ancient, Fe-S scaffold IscA reveals a novel protein fold.
Bilder, P.W., Ding, H., Newcomer, M.E.(2004) Biochemistry 43: 133-139
- PubMed: 14705938
- DOI: https://doi.org/10.1021/bi035440s
- Primary Citation Related Structures:
1R94, 1R95 - PubMed Abstract:
IscA belongs to an ancient family of proteins responsible for iron-sulfur cluster assembly in essential metabolic pathways preserved throughout evolution. We report here the 2.3 A resolution crystal structure of Escherichia coli IscA, a novel fold in which mixed beta-sheets form a compact alpha-beta sandwich domain. In contrast to the highly mobile secondary structural elements within the bacterial Fe-S scaffold protein IscU, a protein which is thought to have a similar function, the great majority of the amino acids that are conserved in IscA homologues are located in elements that constitute a well-ordered fold. However, the 10-residue C-terminal tail segment that contains two invariant cysteines critical for the Fe-S-binding function of a cyanobacterial (Synechocystis PCC) IscA homologue is not ordered in our structure. In addition, the crystal packing reveals a helical assembly that is constructed from two possible tetrameric oligomers of IscA.
- Department of Biological Sciences, Louisiana State University, Baton Rouge, LA 70803, USA.
Organizational Affiliation: