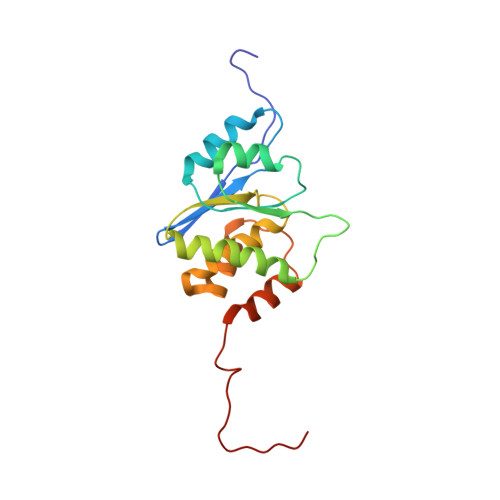
Structural Insights into Molecular Function of the Metastasis-associated Phosphatase PRL-3.
Kozlov, G., Cheng, J., Ziomek, E., Banville, D., Gehring, K., Ekiel, I.(2004) J Biological Chem 279: 11882-11889
- PubMed: 14704153 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M312905200
- Primary Citation Related Structures:
1R6H - PubMed Abstract:
Phosphatases and kinases are the cellular signal transduction enzymes that control protein phosphorylation. PRL phosphatases constitute a novel class of small (20 kDa), prenylated phosphatases with oncogenic activity. In particular, PRL-3 is consistently overexpressed in liver metastasis in colorectal cancer cells and represents a new therapeutic target. Here, we present the solution structure of PRL-3, the first structure of a PRL phosphatase. The structure places PRL phosphatases in the class of dual specificity phosphatases with closest structural homology to the VHR phosphatase. The structure, coupled with kinetic studies of site-directed mutants, identifies functionally important residues and reveals unique features, differentiating PRLs from other phosphatases. These differences include an unusually hydrophobic active site without the catalytically important serine/threonine found in most other phosphatases. The position of the general acid loop indicates the presence of conformational change upon catalysis. The studies also identify a potential regulatory role of Cys(49) that forms an intramolecular disulfide bond with the catalytic Cys(104) even under mildly reducing conditions. Molecular modeling of the highly homologous PRL-1 and PRL-2 phosphatases revealed unique surface elements that are potentially important for specificity.
- Department of Biochemistry, McGill University, Montreal, Quebec H3G 1Y6, Canada.
Organizational Affiliation: