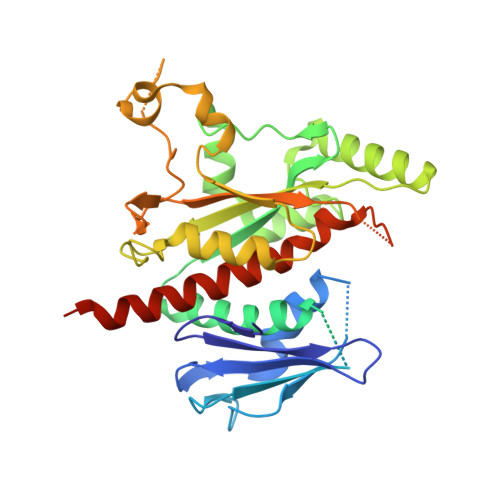
Crystal structure of the bifunctional chorismate synthase from Saccharomyces cerevisiae
Quevillon-Cheruel, S., Leulliot, N., Meyer, P., Graille, M., Bremang, M., Blondeau, K., Sorel, I., Poupon, A., Janin, J., van Tilbeurgh, H.(2004) J Biological Chem 279: 619-625
- PubMed: 14573601 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M310380200
- Primary Citation Related Structures:
1R52, 1R53 - PubMed Abstract:
Chorismate synthase (EC 4.2.3.5), the seventh enzyme in the shikimate pathway, catalyzes the transformation of 5-enolpyruvylshikimate 3-phosphate (EPSP) to chorismate, which is the last common precursor in the biosynthesis of numerous aromatic compounds in bacteria, fungi, and plants. The chorismate synthase reaction involves a 1,4-trans-elimination of phosphoric acid from EPSP and has an absolute requirement for reduced FMN as a cofactor. We have determined the three-dimensional x-ray structure of the yeast chorismate synthase from selenomethionine-labeled crystals at 2.2-A resolution. The structure shows a novel betaalphabetaalpha fold consisting of an alternate tight packing of two alpha-helical and two beta-sheet layers, showing no resemblance to any documented protein structure. The molecule is arranged as a tight tetramer with D2 symmetry, in accordance with its quaternary structure in solution. Electron density is missing for 23% of the amino acids, spread over sequence regions that in the three-dimensional structure converge on the surface of the protein. Many totally conserved residues are contained within these regions, and they probably form a structured but mobile domain that closes over a cleft upon substrate binding and catalysis. This hypothesis is supported by previously published spectroscopic measurements implying that the enzyme undergoes considerable structural changes upon binding of both FMN and EPSP.
- Institut de Biochimie et de Biophysique Moléculaire et Cellulaire (CNRS-UMR 8619), Université Paris-Sud, Bâtiment 430, 91405 Orsay, France.
Organizational Affiliation: