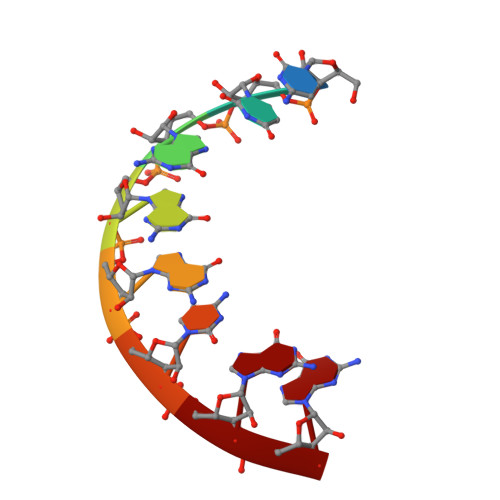
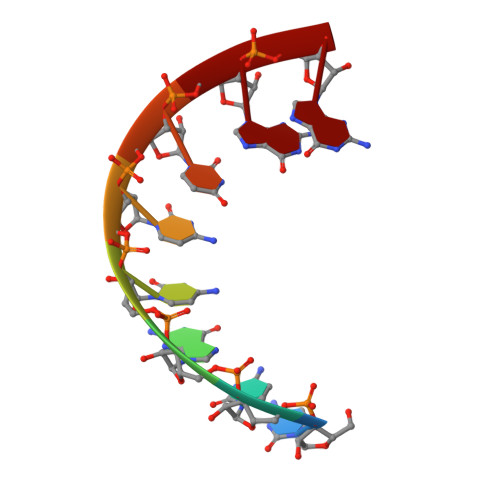
First look at RNA in L-configuration.
Vallazza, M., Perbandt, M., Klussmann, S., Rypniewski, W., Einspahr, H.M., Erdmann, V.A., Betzel, C.h.(2004) Acta Crystallogr D Biol Crystallogr 60: 1-7
- PubMed: 14684885 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444903027690
- Primary Citation Related Structures:
1R3O - PubMed Abstract:
Nucleic acid molecules in the mirror image or L-configuration are unknown in nature and are extraordinarily resistant to biological degradation. The identification of functional L-oligonucleotides called Spiegelmers offers a novel approach for drug discovery based on RNA. The sequence r(CUGGGCGG).r(CCGCCUGG) was chosen as a model system for structural analysis of helices in the L-configuration as the structure of the D-form of this sequence has previously been determined in structural studies of 5S RNA domains, in particular domain E of the Thermus flavus 5S rRNA [Perbandt et al. (2001), Acta Cryst. D57, 219-224]. Unexpectedly, the results of crystallization trials showed little similarity between the D- and the L-forms of the duplex in either the crystallization hits or the diffraction performance. The crystal structure of this L-RNA duplex has been determined at 1.9 A resolution with R(work) and R(free) of 23.8 and 28.6%, respectively. The crystals belong to space group R32, with unit-cell parameters a = 45.7, c = 264.6 A. Although there are two molecules in the asymmetric unit rather than one, the structure of the L-form arranges helical pairs in a head-to-tail fashion to form pseudo-continuous infinite helices in the crystal as in the D-form. On the other hand, the wobble-like G.C(+) base pair seen in the D-RNA analogue does not appear in the L-RNA duplex, which forms a regular double-helical structure with typical Watson-Crick base pairing.
- Institutes of Biology, Chemistry and Pharmacy, Free University of Berlin, Biochemistry: Thielallee 63, 14195 Berlin, Germany.
Organizational Affiliation: