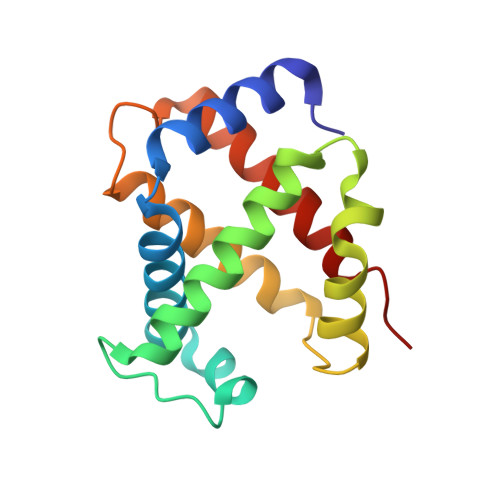
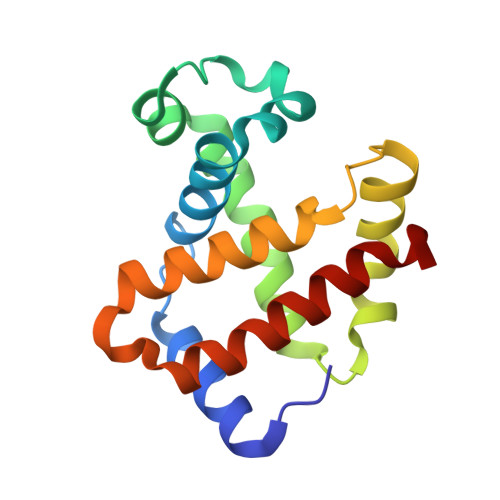
Characterization of hemoglobin bassett (alpha94Asp-->Ala), a variant with very low oxygen affinity
Abdulmalik, O., Safo, M.K., Lerner, N.B., Ochotorena, J., Daikhin, Y., Lakka, V., Santacroce, R., Abraham, D.J., Asakura, T.(2004) Am J Hematol 77: 268-276
- PubMed: 15495251 Search on PubMed
- DOI: https://doi.org/10.1002/ajh.20184
- Primary Citation Related Structures:
1R1X, 1R1Y - PubMed Abstract:
Hemoglobin (Hb) Bassett, an abnormal Hb variant with a markedly reduced oxygen affinity, was discovered in a Caucasian (Anglo-Saxon) male child who experienced episodes of cyanosis. Cation-exchange and reversed-phase (RP) high-performance liquid chromatography (HPLC) showed that the patient has an abnormal Hb, with a mutation in the alpha-globin. Tryptic peptide digest of the abnormal alpha-globin with subsequent HPLC analysis revealed abnormal elution of the alpha-T11 peptide. Further studies with Edman sequencing and electrospray mass spectrometry of tryptic peptide alpha-T11, as well as structural analysis by X-ray crystallography revealed an Asp-->Ala substitution at the alpha94 (G1) position, a match for Hb Bassett. Detailed functional studies showed that this Hb variant had a markedly reduced oxygen affinity (P(50) at pH 7.0 = 22 mmHg; Hb A P(50) = 10.5 mmHg), reduced Bohr effect (-0.26 compared to - 0.54 in Hb A), and low subunit cooperativity (n = 1.4, compared to 2.6 in Hb A). X-ray crystallography results explain the probable effects of the structural modification on the oxygen-binding properties of this Hb variant.
- Division of Hematology, The Children's Hospital of Philadelphia, Philadelphia, Pennsylvania 19104, USA.
Organizational Affiliation: