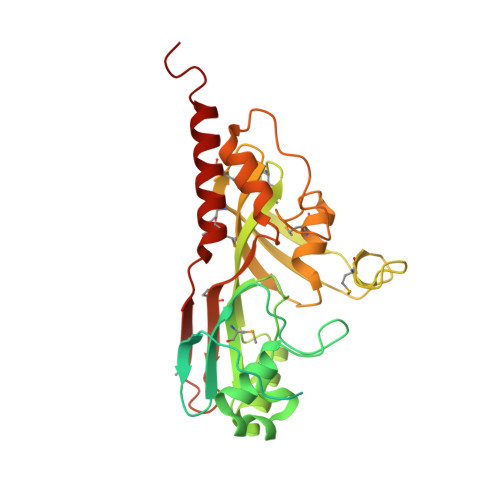
Crystal structure of the catalytic domain of RluD, the only rRNA pseudouridine synthase required for normal growth of Escherichia coli
Del Campo, M., Ofengand, J., Malhotra, A.(2004) RNA 10: 231-239
- PubMed: 14730022 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.5187404
- Primary Citation Related Structures:
1QYU - PubMed Abstract:
Escherichia coli pseudouridine synthase RluD makes pseudouridines 1911, 1915, and 1917 in the loop of helix 69 in 23S RNA. These are the most highly conserved ribosomal pseudouridines known. Of 11 pseudouridine synthases in E. coli, only cells lacking RluD have severe growth defects and abnormal ribosomes. We have determined the 2.0 A structure of the catalytic domain of RluD (residues 77-326), the first structure of an RluA family member. The catalytic domain folds into a mainly antiparallel beta-sheet flanked by several loops and helices. A positively charged cleft that presumably binds RNA leads to the conserved Asp 139. The RluD N-terminal S4 domain, connected by a flexible linker, is disordered in our structure. RluD is very similar in both catalytic domain structure and active site arrangement to the pseudouridine synthases RsuA, TruB, and TruA. We identify five sequence motifs, two of which are novel, in the RluA, RsuA, TruB, and TruA families, uniting them as one superfamily. These results strongly suggest that four of the five families of pseudouridine synthases arose by divergent evolution. The RluD structure also provides insight into its multisite specificity.
- Department of Biochemistry and Molecular Biology, University of Miami School of Medicine, Miami, Florida 33101-6129, USA.
Organizational Affiliation: