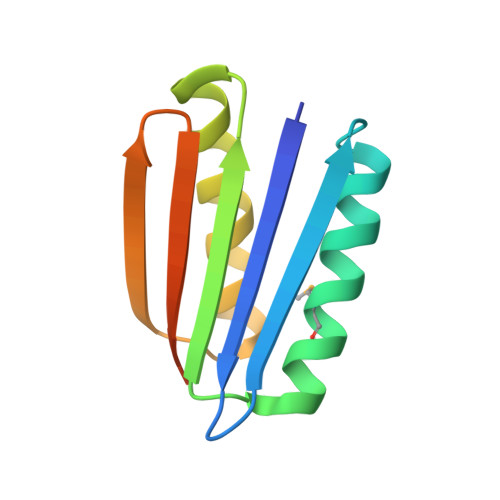
Design of a Novel Globular Protein Fold with Atomic-Level Accuracy
Kuhlman, B., Dantas, G., Ireton, G.C., Varani, G., Stoddard, B.L., Baker, D.(2003) Science 302: 1364-1368
- PubMed: 14631033
- DOI: https://doi.org/10.1126/science.1089427
- Primary Citation Related Structures:
1QYS - PubMed Abstract:
A major challenge of computational protein design is the creation of novel proteins with arbitrarily chosen three-dimensional structures. Here, we used a general computational strategy that iterates between sequence design and structure prediction to design a 93-residue alpha/beta protein called Top7 with a novel sequence and topology. Top7 was found experimentally to be folded and extremely stable, and the x-ray crystal structure of Top7 is similar (root mean square deviation equals 1.2 angstroms) to the design model. The ability to design a new protein fold makes possible the exploration of the large regions of the protein universe not yet observed in nature.
- Department of Biochemistry, University of Washington, Seattle, WA 98195, USA.
Organizational Affiliation: