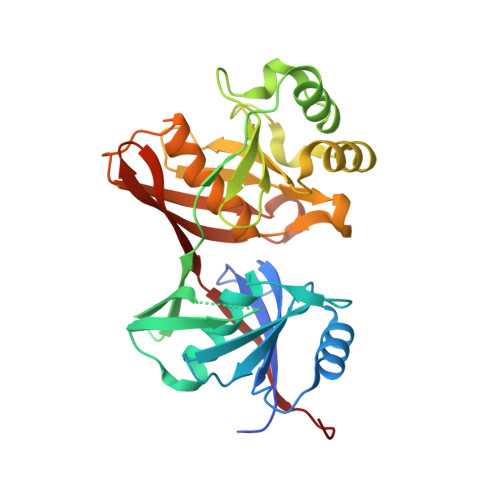
Crystal structure of E. coli yddE protein reveals a striking homology with diaminopimelate epimerase
Grassick, A., Sulzenbacher, G., Roig-Zamboni, V., Campanacci, V., Cambillau, C., Bourne, Y.(2004) Proteins 55: 764-767
- PubMed: 15103639 Search on PubMed
- DOI: https://doi.org/10.1002/prot.20025
- Primary Citation Related Structures:
1QY9, 1QYA - Architecture et Fonction des Macromolécules Biologiques, CNRS UMR6098, Marseille, France.
Organizational Affiliation: