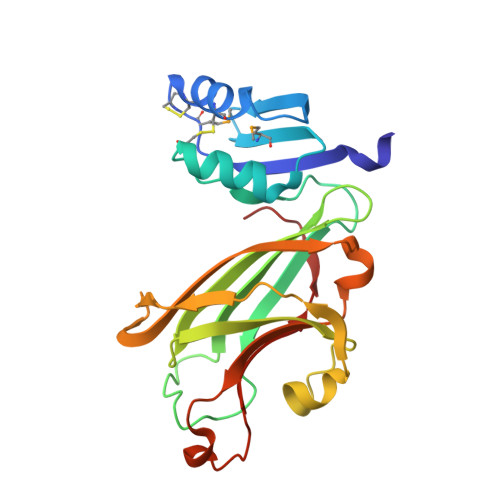
Crystal structure of the copper chaperone for superoxide dismutase.
Lamb, A.L., Wernimont, A.K., Pufahl, R.A., Culotta, V.C., O'Halloran, T.V., Rosenzweig, A.C.(1999) Nat Struct Biol 6: 724-729
- PubMed: 10426947 Search on PubMed
- DOI: https://doi.org/10.1038/11489
- Primary Citation Related Structures:
1QUP - PubMed Abstract:
Cellular systems for handling transition metal ions have been identified, but little is known about the structure and function of the specific trafficking proteins. The 1.8 A resolution structure of the yeast copper chaperone for superoxide dismutase (yCCS) reveals a protein composed of two domains. The N-terminal domain is very similar to the metallochaperone protein Atx1 and is likely to play a role in copper delivery and/or uptake. The second domain resembles the physiological target of yCCS, superoxide dismutase I (SOD1), in overall fold, but lacks all of the structural elements involved in catalysis. In the crystal, two SOD1-like domains interact to form a dimer. The subunit interface is remarkably similar to that in SOD1, suggesting a structural basis for target recognition by this metallochaperone.
- [1] Department of Biochemistry, Molecular Biology, and Cell Biology, Northwestern University, Evanston, Illinois, 60208, USA.
Organizational Affiliation: