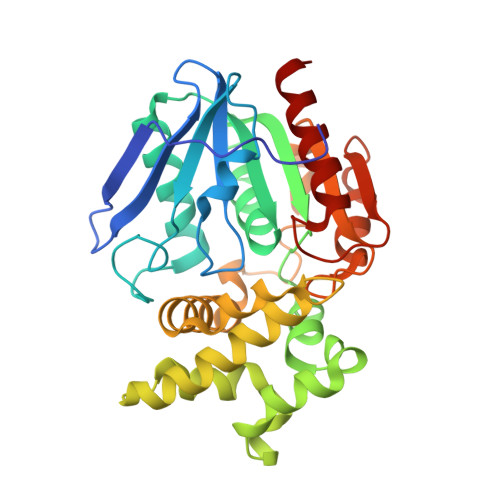
Crystal structure of prolyl aminopeptidase from Serratia marcescens.
Yoshimoto, T., Kabashima, T., Uchikawa, K., Inoue, T., Tanaka, N., Nakamura, K.T., Tsuru, M., Ito, K.(1999) J Biochem 126: 559-565
- PubMed: 10467172
- DOI: https://doi.org/10.1093/oxfordjournals.jbchem.a022486
- Primary Citation of Related Structures:
1QTR - PubMed Abstract:
Prolyl aminopeptidase from Serratia marcescens specifically catalyzes the removal of N-terminal proline residues from peptides. We have solved its three-dimensional structure at 2.3 A resolution by the multiple isomorphous replacement method. The enzyme consists of two contiguous domains. The larger domain shows the general topology of the alpha/beta hydrolase fold, with a central eight-stranded beta-sheet and six helices. The smaller domain consists of six helices. The catalytic triad (Ser113, His296, and Asp268) is located near the large cavity at the interface between the two domains. Cys271, which is sensitive to SH reagents, is located near the catalytic residues, in spite of the fact that the enzyme is a serine peptidase. The specific residues which make up the hydrophobic pocket line the smaller domain, and the specificity of the exo-type enzyme originates from this smaller domain, which blocks the N-terminal of P1 proline.
- School of Pharmaceutical Sciences, Nagasaki University, Nagasaki, 852-8521, Japan. t-yoshimoto@cc.nagasaki-u.ac.jp
Organizational Affiliation: