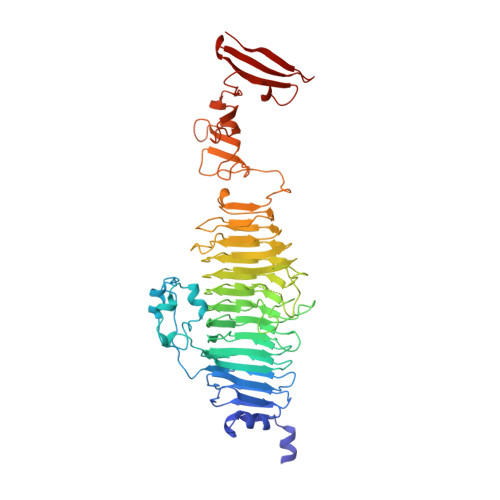
Plasticity and steric strain in a parallel beta-helix: rational mutations in the P22 tailspike protein.
Schuler, B., Furst, F., Osterroth, F., Steinbacher, S., Huber, R., Seckler, R.(2000) Proteins 39: 89-101
- PubMed: 10737931 Search on PubMed
- Primary Citation Related Structures:
1QQ1, 1QRB, 1QRC - PubMed Abstract:
By means of genetic screens, a great number of mutations that affect the folding and stability of the tailspike protein from Salmonella phage P22 have been identified. Temperature-sensitive folding (tsf) mutations decrease folding yields at high temperature, but hardly affect thermal stability of the native trimeric structure when assembled at low temperature. Global suppressor (su) mutations mitigate this phenotype. Virtually all of these mutations are located in the central domain of tailspike, a large parallel beta-helix. We modified tailspike by rational single amino acid replacements at three sites in order to investigate the influence of mutations of two types: (1) mutations expected to cause a tsf phenotype by increasing the side-chain volume of a core residue, and (2) mutations in a similar structural context as two of the four known su mutations, which have been suggested to stabilize folding intermediates and the native structure by the release of backbone strain, an effect well known for residues that are primarily evolved for function and not for stability or folding of the protein. Analysis of folding yields, refolding kinetics and thermal denaturation kinetics in vitro show that the tsf phenotype can indeed be produced rationally by increasing the volume of side chains in the beta-helix core. The high-resolution crystal structure of mutant T326F proves that structural rearrangements only take place in the remarkably plastic lumen of the beta-helix, leaving the arrangement of the hydrogen-bonded backbone and thus the surface of the protein unaffected. This supports the notion that changes in the stability of an intermediate, in which the beta-helix domain is largely formed, are the essential mechanism by which tsf mutations affect tailspike folding. A rational design of su mutants, on the other hand, appears to be more difficult. The exchange of two residues in the active site expected to lead to a drastic release of steric strain neither enhanced the folding properties nor the stability of tailspike. Apparently, side-chain interactions in these cases overcompensate for backbone strain, illustrating the extreme optimization of the tailspike protein for conformational stability. The result exemplifies the view arising from the statistical analysis of the distribution of backbone dihedral angles in known three-dimensional protein structures that the adoption of straight phi/psi angles other than the most favorable ones is often caused by side-chain interactions. Proteins 2000;39:89-101.
- Institut für Biophysik und Physikalische Biochemie, Universität Regensburg, Regensburg, Germany.
Organizational Affiliation: