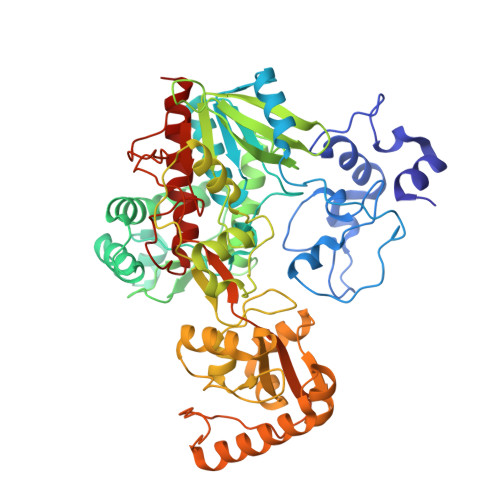
Open Conformation of a Flavocytochrome C3 Fumarate Reductase.
Bamford, V., Dobbin, P.S., Richardson, D.J., Hemmings, A.M.(1999) Nat Struct Biol 6: 1104
- PubMed: 10581549 Search on PubMed
- DOI: https://doi.org/10.1038/70039
- Primary Citation Related Structures:
1QO8 - PubMed Abstract:
Fumarate reductases and succinate dehydrogenases play central roles in the metabolism of eukaryotic and prokaryotic cells. A recent medium resolution structure of the Escherichia coli fumarate reductase (Frd) has revealed the overall organization of the membrane-bound complex. Here we present the first high resolution X-ray crystal structure of a water-soluble bacterial fumarate reductase in an open conformation. This structure reveals a mobile domain that modulates substrate access to the active site and provides new insights into the mechanism of this widespread and important family of FAD-containing respiratory proteins.
- School of Biological Sciences, Centre for Metalloprotein Spectroscopy and Biology, University of East Anglia, Norwich NR4 7TJ, UK.
Organizational Affiliation: