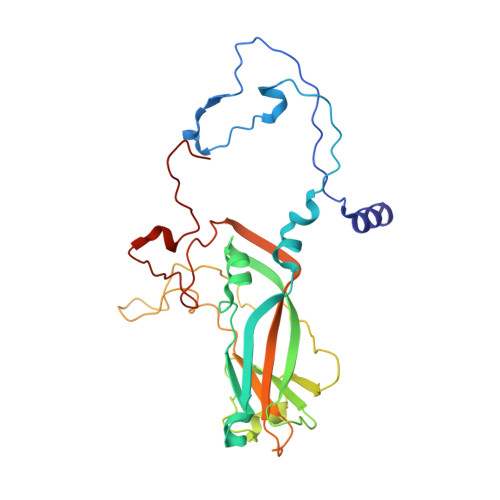
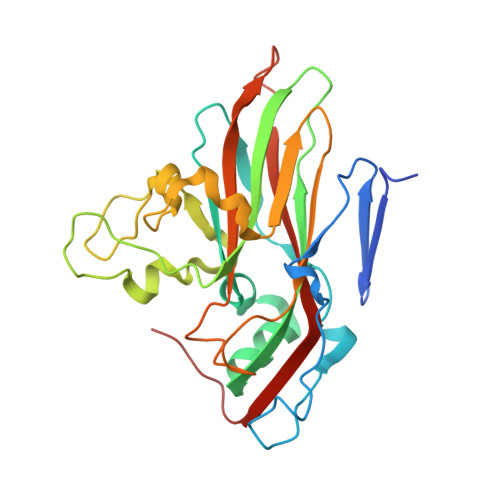
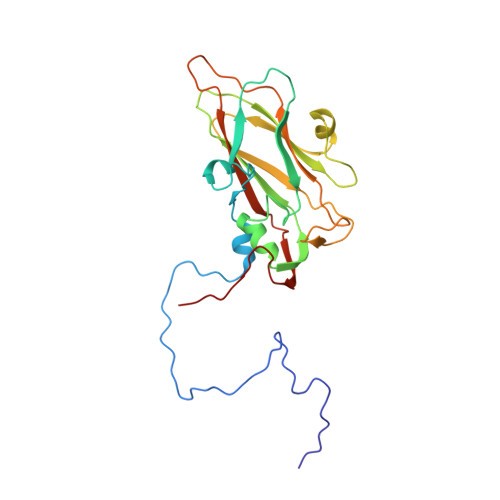
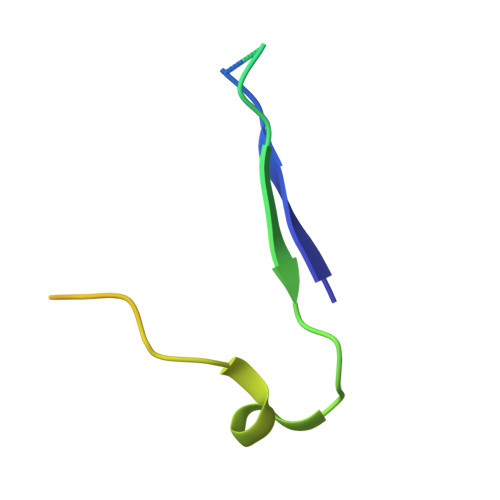
Analysis of Three Structurally Related Antiviral Compounds in Complex with Human Rhinovirus 16
Hadfield, A.T., Minor, I., Diana, G.D., Rossmann, M.G.(1999) Proc Natl Acad Sci U S A 96: 14730
- PubMed: 10611281 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.96.26.14730
- Primary Citation Related Structures:
1QJU, 1QJX, 1QJY - PubMed Abstract:
Rhinoviruses are a frequent cause of the common cold. A series of antirhinoviral compounds have been developed that bind into a hydrophobic pocket in the viral capsid, stabilizing the capsid and interfering with cell attachment. The structures of a variety of such compounds, complexed with rhinovirus serotypes 14, 16, 1A, and 3, previously have been examined. Three chemically similar compounds, closely related to a drug that is undergoing phase III clinical trials, were chosen to determine the structural impact of the heteroatoms in one of the three rings. The compounds were found to have binding modes that depend on their electronic distribution. In the compound with the lowest efficacy, the terminal ring is displaced by 1 A and rotated by 180 degrees relative to the structure of the other two. The greater polarity of the terminal ring in one of the three compounds leads to a small displacement of its position relative to the other compounds in the hydrophobic end of the antiviral compound binding pocket to a site where it makes fewer interactions. Its lower efficacy is likely to be the result of the reduced number of interactions. A region of conserved residues has been identified near the entrance to the binding pocket where there is a corresponding conservation of the mode of binding of these compounds to different serotypes. Thus, variations in residues lining the more hydrophobic end of the pocket are primarily responsible for the differences in drug efficacies.
- Department of Biological Sciences, Purdue University, West Lafayette, IN 47907, USA.
Organizational Affiliation: