Structural Consequences of the B5 Histidine --> Tyrosine Mutation in Human Insulin Characterized by X-Ray Crystallography and Conformational Analysis.
Tang, L., Whittingham, J.L., Verma, C.S., Caves, L.S.D., Dodson, G.G.(1999) Biochemistry 38: 12041
- PubMed: 10508408 Search on PubMed
- DOI: https://doi.org/10.1021/bi990700k
- Primary Citation Related Structures:
1QIY, 1QIZ, 1QJ0 - PubMed Abstract:
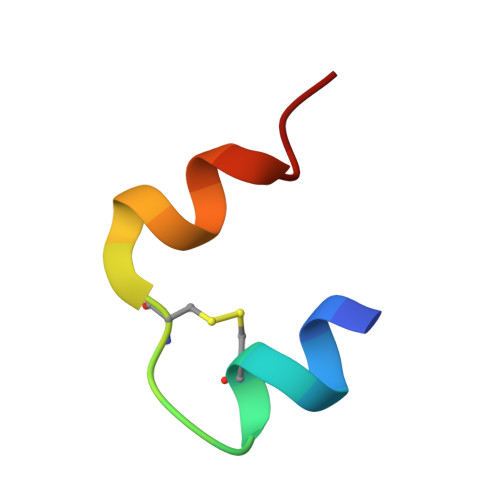
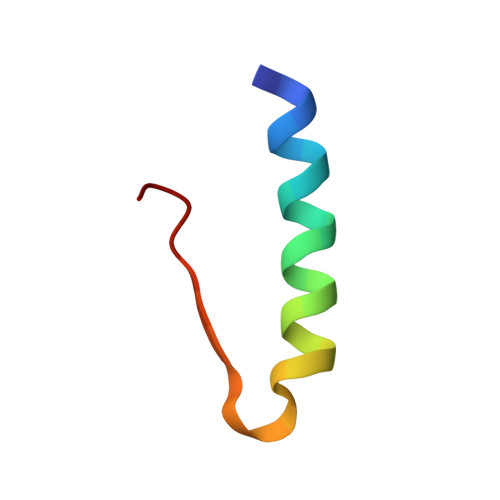
The addition of phenols to hexameric insulin solutions produces a particularly stable hexamer, resulting from a rearrangement in which residues B1-B8 change from an extended conformation (T-state) to form an alpha-helix (R-state). The R-state is, in part, stabilized by nonpolar interactions between the phenolic molecule and residue B5 His at the dimer-dimer interface. The B5 His --> Tyr mutant human insulin was constructed to see if the tyrosine side chain would mimic the effect of phenol binding in the hexamer and induce the R-state. In partial support of this hypothesis, the molecule crystallized as a half-helical hexamer (T(3)R(3)) in conditions that conventionally promote the fully nonhelical (T6) form. As expected, in the presence of phenol or resorcinol, the B5 Tyr hexamers adopt the fully helical (R6) conformation. Molecular modeling calculations were performed to investigate the conformational preference of the T-state B5 Tyr side chain in the T(3)R(3) form, this side chain being associated with structural perturbations of the A7-A10 loop in an adjacent hexamer. For an isolated dimer, several different orientations of the side chain were found, which were close in energy and readily interconvertible. In the crystal environment only one of these conformations remains low in energy; this conformation corresponds to that observed in the crystal structure. This suggests that packing constraints around residue B5 Tyr result in the observed structural rearrangements. Thus, rather than promoting the R-state in a manner analogous to phenol, the mutation appears to destabilize the T-state. These studies highlight the role of B5 His in determining hexamer conformation and in mediating crystal packing interactions, properties that are likely be important in vivo.
- Protein Structure Division, National Institute for Medical Research, London, England.
Organizational Affiliation: