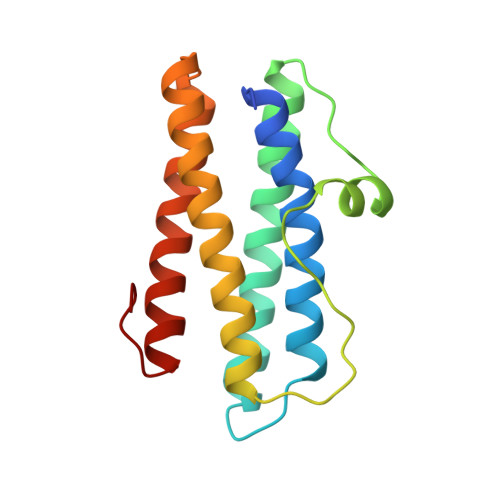
The dodecameric ferritin from Listeria innocua contains a novel intersubunit iron-binding site.
Ilari, A., Stefanini, S., Chiancone, E., Tsernoglou, D.(2000) Nat Struct Biol 7: 38-43
- PubMed: 10625425 Search on PubMed
- DOI: https://doi.org/10.1038/71236
- Primary Citation Related Structures:
1QGH - PubMed Abstract:
Ferritin is characterized by a highly conserved architecture that comprises 24 subunits assembled into a spherical cage with 432 symmetry. The only known exception is the dodecameric ferritin from Listeria innocua. The structure of Listeria ferritin has been determined to a resolution of 2.35 A by molecular replacement, using as a search model the structure of Dps from Escherichia coli. The Listeria 12-mer is endowed with 23 symmetry and displays the functionally relevant structural features of the ferritin 24-mer, namely the negatively charged channels along the three-fold symmetry axes that serve for iron entry into the cavity and a negatively charged internal cavity for iron deposition. The electron density map shows 12 iron ions on the inner surface of the hollow core, at the interface between monomers related by two-fold axes. Analysis of the nature and stereochemistry of the iron-binding ligands reveals strong similarities with known ferroxidase sites. The L. innocua ferritin site, however, is the first described so far that has ligands belonging to two different subunits and is not contained within a four-helix bundle.
- CNR Center of Molecular Biology, Department of Biochemical Sciences "A. Rossi Fanelli", University La Sapienza, Piazzale Aldo Moro 5, 00185 Rome, Italy.
Organizational Affiliation: