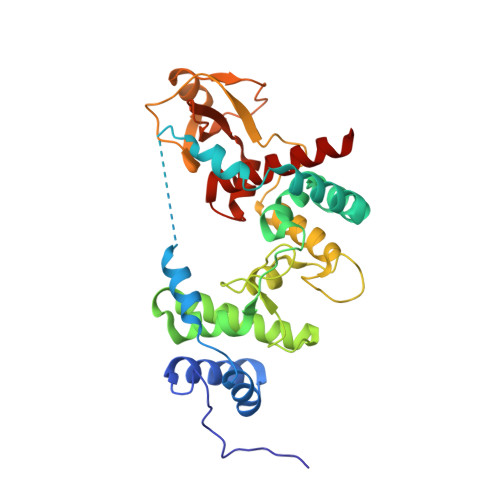
Binding of calcium in the EF-hand of Escherichia coli lytic transglycosylase Slt35 is important for stability.
van Asselt, E.J., Dijkstra, B.W.(1999) FEBS Lett 458: 429-435
- PubMed: 10570954 Search on PubMed
- DOI: https://doi.org/10.1016/s0014-5793(99)01198-9
- Primary Citation Related Structures:
1QDR, 1QDT - PubMed Abstract:
The Escherichia coli lytic transglycosylase Slt35 contains a single metal ion-binding site that resembles EF-hand calcium-binding sites. The Slt35 EF-hand is only the second observation of such a domain in a prokaryotic protein. Two crystal structures at 2.1 A resolution show that both Ca2+ ions and Na+ ions can bind to the EF-hand domain, but in subtly different configurations. Heat-induced unfolding studies demonstrate that Ca2+ ions are preferentially bound, and that only Ca2+ ions significantly increase the melting temperature of Slt35. This shows that the EF-hand calcium-binding domain is important for the stability of Slt35.
- BIOSON Research Institute and Laboratory of Biophysical Chemistry, University of Groningen, The Netherlands.
Organizational Affiliation: