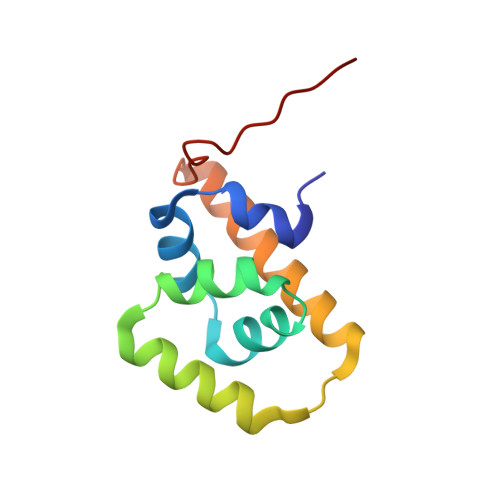
Structure of the C-terminal domain of FliG, a component of the rotor in the bacterial flagellar motor
Lloyd, S.A., Whitby, F.G., Blair, D.F., Hill, C.P.(1999) Nature 400: 472-475
- PubMed: 10440379 Search on PubMed
- DOI: https://doi.org/10.1038/22794
- Primary Citation Related Structures:
1QC7 - PubMed Abstract:
Many motile species of bacteria are propelled by flagella, which are rigid helical filaments turned by rotary motors in the cell membrane. The motors are powered by the transmembrane gradient of protons or sodium ions. Although bacterial flagella contain many proteins, only three-MotA, MotB and FliG-participate closely in torque generation. MotA and MotB are ion-conducting membrane proteins that form the stator of the motor. FliG is a component of the rotor, present in about 25 copies per flagellum. It is composed of an amino-terminal domain that functions in flagellar assembly and a carboxy-terminal domain (FliG-C) that functions specifically in motor rotation. Here we report the crystal structure of FliG-C from the hyperthermophilic eubacterium Thermotoga maritima. Charged residues that are important for function, and which interact with the stator protein MotA, cluster along a prominent ridge on FliG-C. On the basis of the disposition of these residues, we present a hypothesis for the orientation of FliG-C domains in the flagellar motor, and propose a structural model for the part of the rotor that interacts with the stator.
- Department of Biology, University of Utah, Salt Lake City 84112-0840, USA.
Organizational Affiliation: