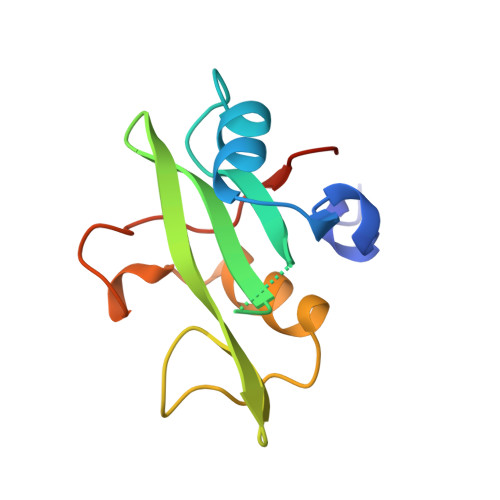
Crystal structure of the C-terminal SH2 domain of the p85alpha regulatory subunit of phosphoinositide 3-kinase: an SH2 domain mimicking its own substrate.
Hoedemaeker, F.J., Siegal, G., Roe, S.M., Driscoll, P.C., Abrahams, J.P.(1999) J Mol Biology 292: 763-770
- PubMed: 10525402 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.3111
- Primary Citation Related Structures:
1QAD - PubMed Abstract:
The binding properties of Src homology-2 (SH2) domains to phosphotyrosine (pY)-containing peptides have been studied in recent years with the elucidation of a large number of crystal and solution structures. Taken together, these structures suggest a general mode of binding of pY-containing peptides, explain the specificities of different SH2 domains, and may be used to design inhibitors of pY binding by SH2 domain-containing proteins. We now report the crystal structure to 1.8 A resolution of the C-terminal SH2 domain (C-SH2) of the P85alpha regulatory subunit of phosphoinositide 3-kinase (PI3 K). Surprisingly, the carboxylate group of Asp2 from a neighbouring molecule occupies the phosphotyrosine binding site and interacts with Arg18 (alphaA2) and Arg36 (betaB5), in a similar manner to the phosphotyrosine-protein interactions seen in structures of other SH2 domains complexed with pY peptides. It is the first example of a non-phosphate-containing, non-aromatic mimetic of phosphotyrosine binding to SH2 domains, and this could have implications for the design of substrate analogues and inhibitors. Overall, the crystal structure closely resembles the solution structure, but a number of loops which demonstrate mobility in solution are well defined by the crystal packing. C-SH2 has adopted a binding conformation reminiscent of the ligand bound N-terminal SH2 domain of PI3K, apparently induced by the substrate mimicking of a neighbouring molecule in the crystal.
- Leiden Institute for Chemistry Gorlaeus Laboratoria, Universiteit Leiden, 2300 RA, The Netherlands. hoedemae@chem.leidenuniv.nl
Organizational Affiliation: