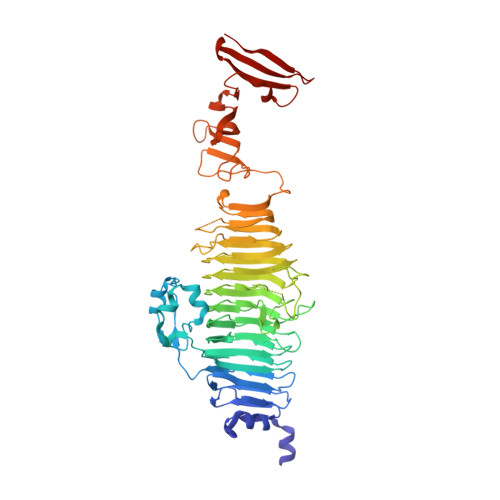
Mutations improving the folding of phage P22 tailspike protein affect its receptor binding activity.
Baxa, U., Steinbacher, S., Weintraub, A., Huber, R., Seckler, R.(1999) J Mol Biology 293: 693-701
- PubMed: 10543960 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.3165
- Primary Citation Related Structures:
1CLW, 1QA1, 1QA2, 1QA3 - PubMed Abstract:
Four previously isolated mutations in Salmonella phage P22 tailspike protein were used to study the relationship between protein stability, folding, and function. Tailspike protein binds and hydrolyzes the repetitive O-antigen structure in Salmonella lipopolysaccharide. Four mutations (V331G, V331A, A334V, A334I) are known to increase the folding efficiency, and two of them (at position 331) also increase the thermal stability of the protein. Octasaccharides comprising two repeating units of the O-antigens from two different Salmonella strains were employed to analyze the receptor binding function of the mutant proteins. Their endorhamnosidase enzymatic activity was assayed with the aid of a fluorescence-labeled dodecasaccharide. Both V331A and V331G were found to strongly affect O-antigen binding. Octasaccharide binding affinities of the mutant proteins are reduced tenfold and 200-fold, corresponding to a loss of 17% and 36% of the standard free energy of binding, respectively. Both mutations at position 334 affected O-antigen binding only slightly (DeltaDeltaG(0)B approximately 1 kJ/mol), but these mutations reduce the thermal stability of the protein. The observed effects on the endoglycosidase activity are fully explained by the changes in substrate binding, suggesting that neither of the mutations affect the catalytic rate. Crystal structures of all four mutants were determined to a resolution of 2.0 A. Except for the partly or completely missing side-chain, no significant changes compared to the wild-type protein structure were found for the mutants at position 331, whereas a small but significant backbone displacement around the mutation site in A334V and A334I may explain the observed thermal destabilization.
- Physikalische Biochemie, Universität Potsdam, Im Biotechnologiepark, Luckenwalde, D-14943, Germany.
Organizational Affiliation: