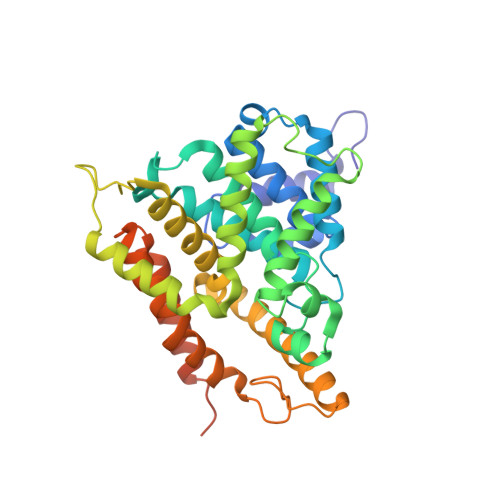
Three dimensional structures of PDE4D in complex with roliprams and implication on inhibitor selectivity
Huai, Q., Wang, H., Sun, Y., Kim, H.Y., Liu, Y., Ke, H.(2003) Structure 11: 865-873
- PubMed: 12842049 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(03)00123-0
- Primary Citation Related Structures:
1OYN, 1Q9M - PubMed Abstract:
Selective inhibitors against the 11 families of cyclic nucleotide phosphodiesterases (PDEs) are used to treat various human diseases. How the inhibitors selectively bind the conserved PDE catalytic domains is unknown. The crystal structures of the PDE4D2 catalytic domain in complex with (R)- or (R,S)-rolipram suggest that inhibitor selectivity is determined by the chemical nature of amino acids and subtle conformational changes of the binding pockets. The conformational states of Gln369 in PDE4D2 may play a key role in inhibitor recognition. The corresponding Y329S mutation in PDE7 may lead to loss of the hydrogen bonds between rolipram and Gln369 and is thus a possible reason explaining PDE7's insensitivity to rolipram inhibition. Docking of the PDE5 inhibitor sildenafil into the PDE4 catalytic pocket further helps understand inhibitor selectivity.
- Department of Biochemistry and Biophysics and Lineberger Comprehensive Cancer Center, The University of North Carolina, Chapel Hill, Chapel Hill, NC 27599, USA.
Organizational Affiliation: