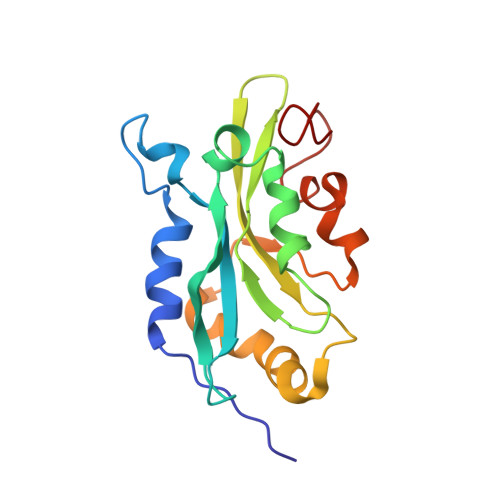
The solution structure of human cofilin: rationalizing actin binding and pH sensitivity
Pope, B.J., Zierler-Gould, K.M., Kuhne, R., Weeds, A.G., Ball, L.J.(2004) J Biological Chem 279: 4840-4848
- PubMed: 14627701
- DOI: https://doi.org/10.1074/jbc.M310148200
- Primary Citation of Related Structures:
1Q8G, 1Q8X - PubMed Abstract:
Human actin-depolymerizing factor (ADF) and cofilin are pH-sensitive, actin-depolymerizing proteins. Although 72% identical in sequence, ADF has a much higher depolymerizing activity than cofilin at pH 8. To understand this, we solved the structure of human cofilin using nuclear magnetic resonance and compared it with human ADF. Important sequence differences between vertebrate ADF/cofilins were correlated with unique structural determinants in the F-actin-binding site to account for differences in biochemical activities of the two proteins. Cofilin has a short beta-strand at the C terminus, not found in ADF, which packs against strands beta3/beta4, changing the environment around Lys96, a residue essential for F-actin binding. A salt bridge involving His133 and Asp98 (Glu98 in ADF) may explain the pH sensitivity of human cofilin and ADF; these two residues are fully conserved in vertebrate ADF/cofilins. Chemical shift perturbations identified residues that (i) differ in their chemical environments between wild type cofilin and mutants S3D, which has greatly reduced G-actin binding, and K96Q, which does not bind F-actin; (ii) are affected when G-actin binds cofilin; and (iii) are affected by pH change from 6 to 8. Many residues affected by G-actin binding also show perturbation in the mutants or in response to pH. Our evidence suggests the involvement of residues 133-138 of strand beta5 in all of the activities examined. Because residues in beta5 are perturbed by mutations that affect both G-actin and F-actin binding, this strand forms a "boundary" or "bridge" between the proposed F- and G-actin-binding sites.
- Medical Research Council Laboratory of Molecular Biology, Hills Road, Cambridge CB2 2QH, United Kingdom.
Organizational Affiliation: